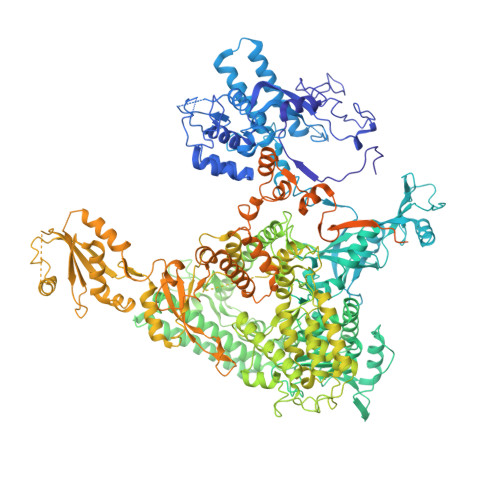
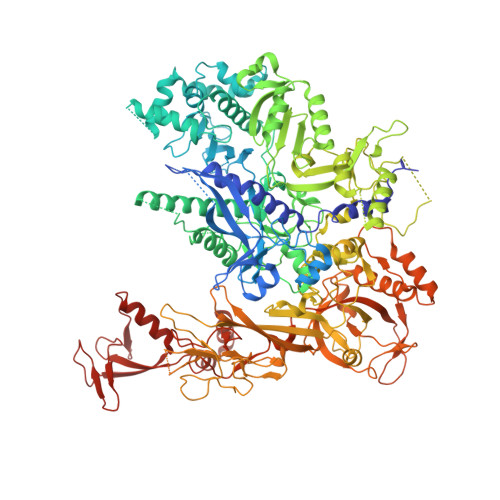
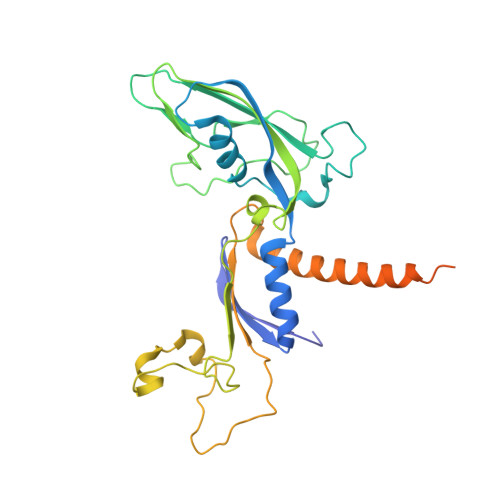
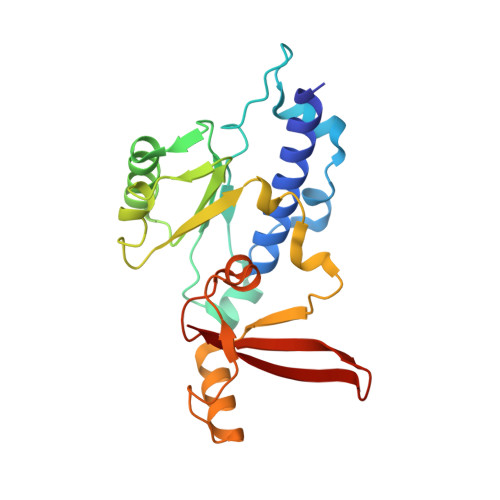
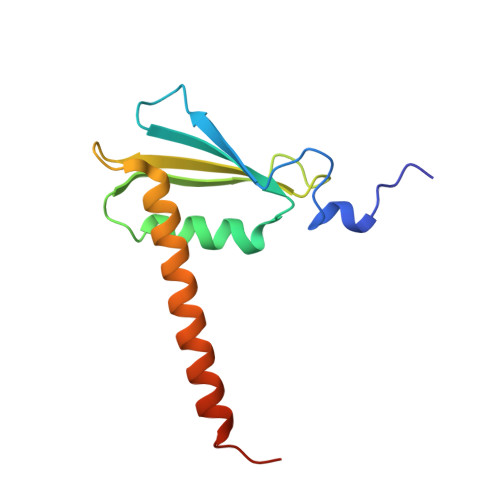
Epigenetic DNA Modification N6-Methyladenine Causes Site-Specific RNA Polymerase II Transcriptional Pausing.
Wang, W., Xu, L., Hu, L., Chong, J., He, C., Wang, D.(2017) J Am Chem Soc 139: 14436-14442
- PubMed: 28933854 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/jacs.7b06381
- Primary Citation Related Structures:
5W4U, 5W51 - PubMed Abstract:
N 6 -Methyladenine (N 6 -mA or 6 mA) is an epigenetic DNA modification in eukaryotic genomes. In contrast to the well-established roles of 5-methylcytosine for epigenetic regulation of gene expression, the functional roles of N 6 -mA remain elusive. In particular, the impact of N 6 -mA modification of the DNA template on RNA polymerase II (pol II) transcription elongation is not known. In this work, using the Saccharomyces cerevisiae pol II transcriptional elongation system as a model, we investigated the molecular mechanism of pol II recognition and processing of N 6 -mA sites via both biochemical and structural approaches. We found that N 6 -mA causes site-specific pol II pausing/stalling. Structural analysis revealed that while N 6 -mA can reach the +1 template position, the stability of the N 6 -mA and UTP base pairing is compromised. Taken together, we reveal that the presence of the 6-methyl group on adenine reduces incorporation efficiency and promotes backtracking translocation. Our studies with yeast pol II provide molecular insights into understanding the impacts of N 6 -mA on pol II transcription dynamics in different organisms.
- Department of Chemistry, Sun Yat-Sen University , Guangzhou 510275, China.
Organizational Affiliation: