Mechanism of Error-Free DNA Replication Past Lucidin-Derived DNA Damage by Human DNA Polymerase kappa.
Yockey, O.P., Jha, V., Ghodke, P.P., Xu, T., Xu, W., Ling, H., Pradeepkumar, P.I., Zhao, L.(2017) Chem Res Toxicol 30: 2023-2032
- PubMed: 28972744 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.chemrestox.7b00227
- Primary Citation Related Structures:
5W2A, 5W2C - PubMed Abstract:
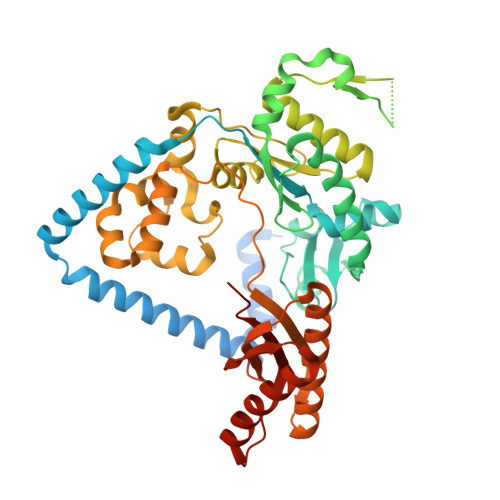
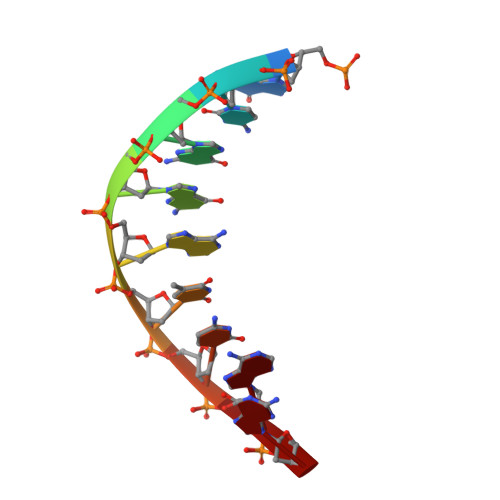
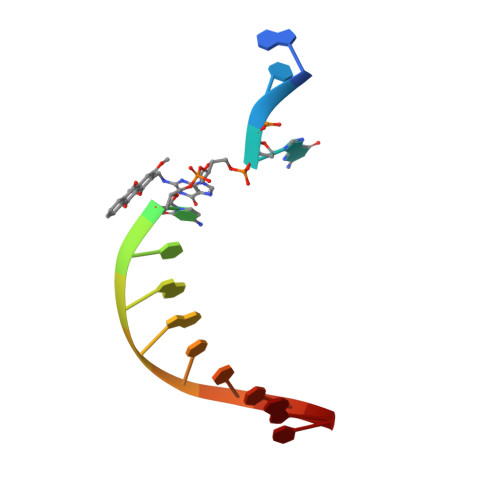
DNA damage impinges on genetic information flow and has significant implications in human disease and aging. Lucidin-3-O-primeveroside (LuP) is an anthraquinone derivative present in madder root, which has been used as a coloring agent and food additive. LuP can be metabolically converted to genotoxic compound lucidin, which subsequently forms lucidin-specific N 2 -2'-deoxyguanosine (N 2 -dG) and N 6 -2'-deoxyadenosine (N 6 -dA) DNA adducts. Lucidin is mutagenic and carcinogenic in rodents but has low carcinogenic risks in humans. To understand the molecular mechanism of low carcinogenicity of lucidin in humans, we performed DNA replication assays using site-specifically modified oligodeoxynucleotides containing a structural analogue (LdG) of lucidin-N 2 -dG DNA adduct and determined the crystal structures of DNA polymerase (pol) κ in complex with LdG-bearing DNA and an incoming nucleotide. We examined four human pols (pol η, pol ι, pol κ, and Rev1) in their efficiency and accuracy during DNA replication with LdG; these pols are key players in translesion DNA synthesis. Our results demonstrate that pol κ efficiently and accurately replicates past the LdG adduct, whereas DNA replication by pol η, pol ι is compromised to different extents. Rev1 retains its ability to incorporate dCTP opposite the lesion albeit with decreased efficiency. Two ternary crystal structures of pol κ illustrate that the LdG adduct is accommodated by pol κ at the enzyme active site during insertion and postlesion-extension steps. The unique open active site of pol κ allows the adducted DNA to adopt a standard B-form for accurate DNA replication. Collectively, these biochemical and structural data provide mechanistic insights into the low carcinogenic risk of lucidin in humans.
- Department of Biochemistry, Schulich School of Medicine & Dentistry, University of Western Ontario , London, Ontario N6A 5C1, Canada.
Organizational Affiliation: