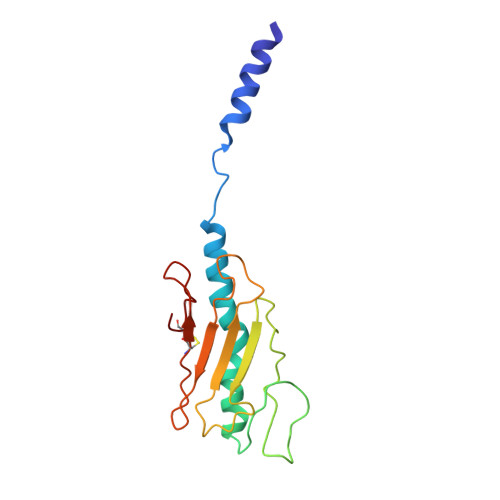
Cryoelectron Microscopy Reconstructions of the Pseudomonas aeruginosa and Neisseria gonorrhoeae Type IV Pili at Sub-nanometer Resolution.
Wang, F., Coureuil, M., Osinski, T., Orlova, A., Altindal, T., Gesbert, G., Nassif, X., Egelman, E.H., Craig, L.(2017) Structure 25: 1423-1435.e4
- PubMed: 28877506 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2017.07.016
- Primary Citation Related Structures:
5VXX, 5VXY - PubMed Abstract:
We report here cryoelectron microscopy reconstructions of type IV pili (T4P) from two important human pathogens, Pseudomonas aeruginosa and Neisseria gonorrhoeae, at ∼ 8 and 5 Å resolution, respectively. The two structures reveal distinct arrangements of the pilin globular domains on the pilus surfaces, which impart different helical parameters, but similar packing of the conserved N-terminal α helices, α1, in the filament core. In contrast to the continuous α helix seen in the X-ray crystal structures of the P. aeruginosa and N. gonorrhoeae pilin subunits, α1 in the pilus filaments has a melted segment located between conserved helix-breaking residues Gly14 and Pro22, as seen for the Neisseria meningitidis T4P. Using mutagenesis we show that Pro22 is critical for pilus assembly, as are Thr2 and Glu5, which are positioned to interact in the hydrophobic filament core. These structures provide a framework for understanding T4P assembly, function, and biophysical properties.
- Department of Biochemistry and Molecular Genetics, University of Virginia School of Medicine, Charlottesville, VA 22908, USA.
Organizational Affiliation: