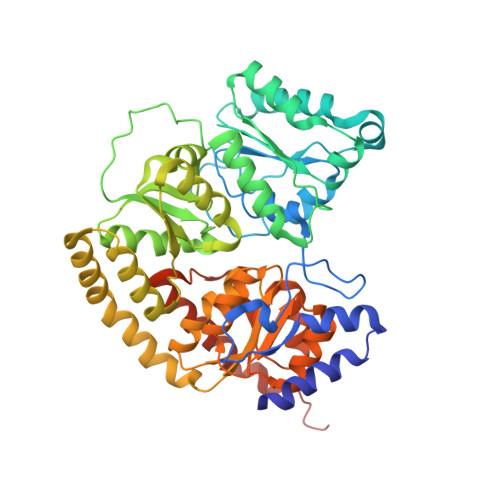
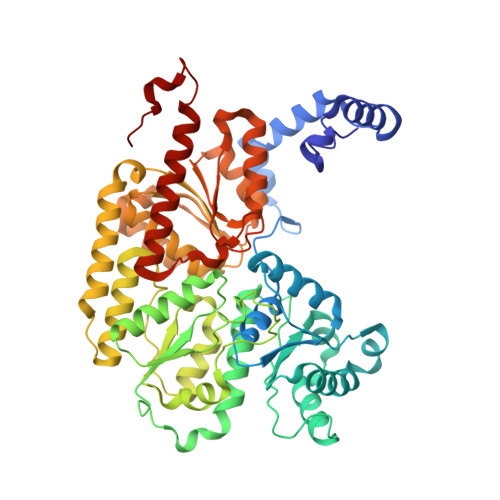
Reversible Protonated Resting State of the Nitrogenase Active Site.
Morrison, C.N., Spatzal, T., Rees, D.C.(2017) J Am Chem Soc 139: 10856-10862
- PubMed: 28692802 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/jacs.7b05695
- Primary Citation Related Structures:
5VPW, 5VQ3, 5VQ4 - PubMed Abstract:
Protonated states of the nitrogenase active site are mechanistically significant since substrate reduction is invariably accompanied by proton uptake. We report the low pH characterization by X-ray crystallography and EPR spectroscopy of the nitrogenase molybdenum iron (MoFe) proteins from two phylogenetically distinct nitrogenases (Azotobacter vinelandii, Av, and Clostridium pasteurianum, Cp) at pHs between 4.5 and 8. X-ray data at pHs of 4.5-6 reveal the repositioning of side chains along one side of the FeMo-cofactor, and the corresponding EPR data shows a new S = 3/2 spin system with spectral features similar to a state previously observed during catalytic turnover. The structural changes suggest that FeMo-cofactor belt sulfurs S3A or S5A are potential protonation sites. Notably, the observed structural and electronic low pH changes are correlated and reversible. The detailed structural rearrangements differ between the two MoFe proteins, which may reflect differences in potential protonation sites at the active site among nitrogenase species. These observations emphasize the benefits of investigating multiple nitrogenase species. Our experimental data suggest that reversible protonation of the resting state is likely occurring, and we term this state "E 0 H + ", following the Lowe-Thorneley naming scheme.
- Division of Chemistry and Chemical Engineering and ‡Howard Hughes Medical Institute, California Institute of Technology , Pasadena, California 91125, United States.
Organizational Affiliation: