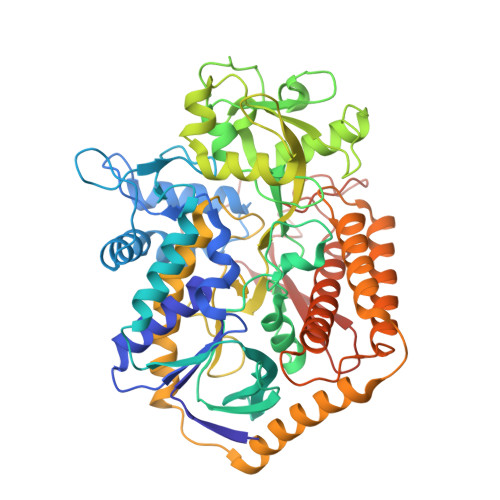
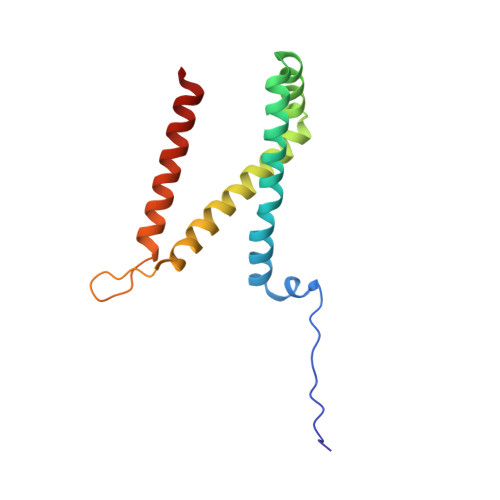
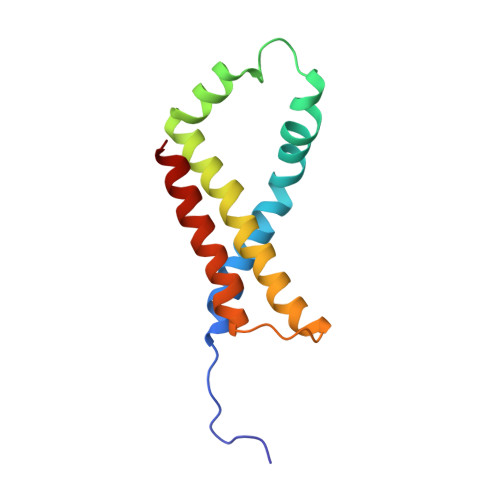
Structural and biochemical analyses reveal insights into covalent flavinylation of the Escherichia coli Complex II homolog quinol:fumarate reductase.
Starbird, C.A., Maklashina, E., Sharma, P., Qualls-Histed, S., Cecchini, G., Iverson, T.M.(2017) J Biological Chem 292: 12921-12933
- PubMed: 28615448 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M117.795120
- Primary Citation Related Structures:
5VPN - PubMed Abstract:
The Escherichia coli Complex II homolog quinol:fumarate reductase (QFR, FrdABCD) catalyzes the interconversion of fumarate and succinate at a covalently attached FAD within the FrdA subunit. The SdhE assembly factor enhances covalent flavinylation of Complex II homologs, but the mechanisms underlying the covalent attachment of FAD remain to be fully elucidated. Here, we explored the mechanisms of covalent flavinylation of the E. coli QFR FrdA subunit. Using a Δ sdhE E. coli strain, we show that the requirement for the assembly factor depends on the cellular redox environment. We next identified residues important for the covalent attachment and selected the FrdA E245 residue, which contributes to proton shuttling during fumarate reduction, for detailed biophysical and structural characterization. We found that QFR complexes containing FrdA E245Q have a structure similar to that of the WT flavoprotein, but lack detectable substrate binding and turnover. In the context of the isolated FrdA subunit, the anticipated assembly intermediate during covalent flavinylation, FrdA E245 variants had stability similar to that of WT FrdA, contained noncovalent FAD, and displayed a reduced capacity to interact with SdhE. However, small-angle X-ray scattering (SAXS) analysis of WT FrdA cross-linked to SdhE suggested that the FrdA E245 residue is unlikely to contribute directly to the FrdA-SdhE protein-protein interface. We also found that no auxiliary factor is absolutely required for flavinylation, indicating that the covalent flavinylation is autocatalytic. We propose that multiple factors, including the SdhE assembly factor and bound dicarboxylates, stimulate covalent flavinylation by preorganizing the active site to stabilize the quinone-methide intermediate.
- Graduate Program in Chemical and Physical Biology, Vanderbilt University, Nashville, Tennessee 37232.
Organizational Affiliation: