Molecular Determinants For Antibody Binding On Group 1 House Dust Mite Allergens.
Chruszcz, M., Pomes, A., Glesner, J., Vailes, L.D., Osinski, T., Porebski, P.J., Majorek, K.A., Heymann, P.W., Platts-Mills, T.A., Minor, W., Chapman, M.D.(2012) J Biological Chem 287: 7388
- PubMed: 22210776 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M111.311159
- Primary Citation Related Structures:
3RVT, 3RVU, 5VPG, 5VPH, 5VPL - PubMed Abstract:
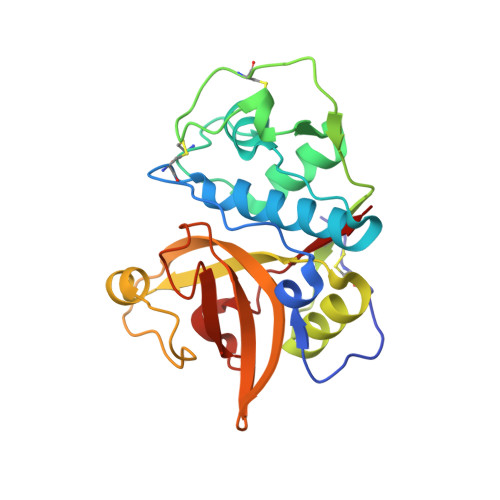
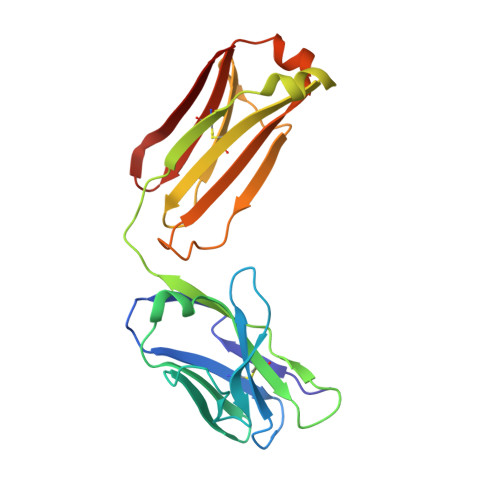
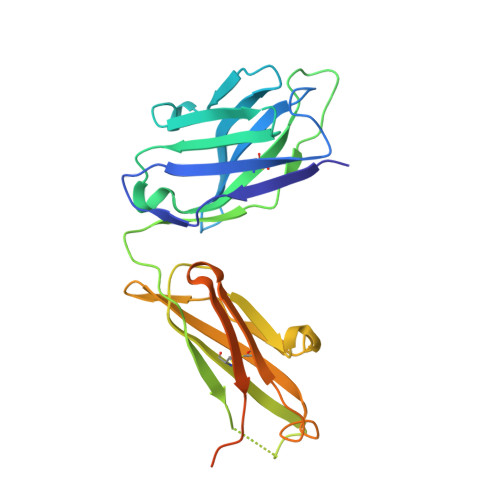
House dust mites produce potent allergens, Der p 1 and Der f 1, that cause allergic sensitization and asthma. Der p 1 and Der f 1 are cysteine proteases that elicit IgE responses in 80% of mite-allergic subjects and have proinflammatory properties. Their antigenic structure is unknown. Here, we present crystal structures of natural Der p 1 and Der f 1 in complex with a monoclonal antibody, 4C1, which binds to a unique cross-reactive epitope on both allergens associated with IgE recognition. The 4C1 epitope is formed by almost identical amino acid sequences and contact residues. Mutations of the contact residues abrogate mAb 4C1 binding and reduce IgE antibody binding. These surface-exposed residues are molecular targets that can be exploited for development of recombinant allergen vaccines.
- Department of Molecular Physiology and Biological Physics, University of Virginia, Charlottesville, Virginia 22908, USA.
Organizational Affiliation: