Recurrent Potent Human Neutralizing Antibodies to Zika Virus in Brazil and Mexico.
Robbiani, D.F., Bozzacco, L., Keeffe, J.R., Khouri, R., Olsen, P.C., Gazumyan, A., Schaefer-Babajew, D., Avila-Rios, S., Nogueira, L., Patel, R., Azzopardi, S.A., Uhl, L.F.K., Saeed, M., Sevilla-Reyes, E.E., Agudelo, M., Yao, K.H., Golijanin, J., Gristick, H.B., Lee, Y.E., Hurley, A., Caskey, M., Pai, J., Oliveira, T., Wunder, E.A., Sacramento, G., Nery, N., Orge, C., Costa, F., Reis, M.G., Thomas, N.M., Eisenreich, T., Weinberger, D.M., de Almeida, A.R.P., West, A.P., Rice, C.M., Bjorkman, P.J., Reyes-Teran, G., Ko, A.I., MacDonald, M.R., Nussenzweig, M.C.(2017) Cell 169: 597-609.e11
- PubMed: 28475892 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.cell.2017.04.024
- Primary Citation Related Structures:
5VIC, 5VIG - PubMed Abstract:
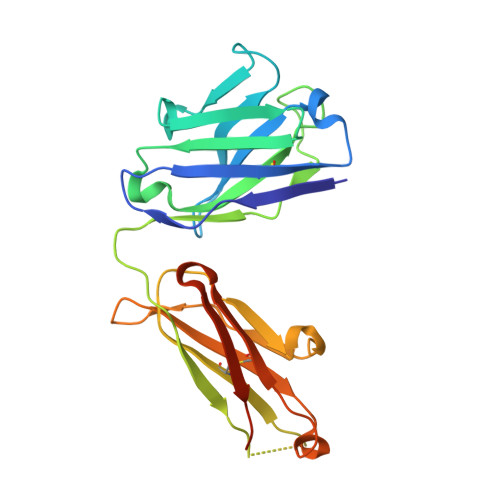
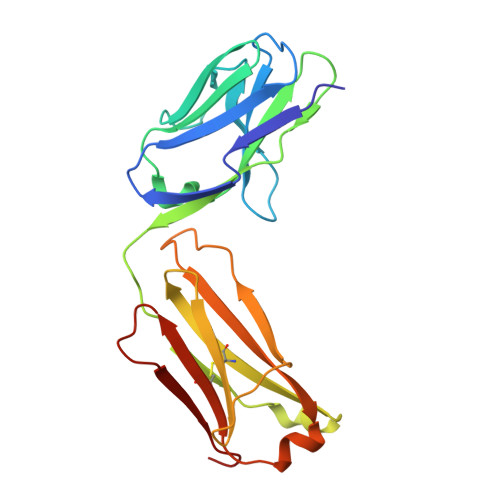
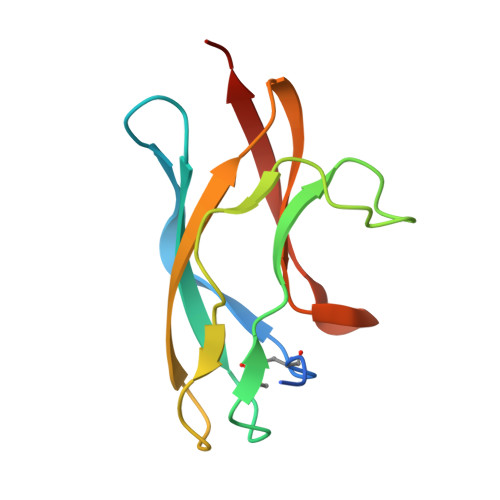
Antibodies to Zika virus (ZIKV) can be protective. To examine the antibody response in individuals who develop high titers of anti-ZIKV antibodies, we screened cohorts in Brazil and Mexico for ZIKV envelope domain III (ZEDIII) binding and neutralization. We find that serologic reactivity to dengue 1 virus (DENV1) EDIII before ZIKV exposure is associated with increased ZIKV neutralizing titers after exposure. Antibody cloning shows that donors with high ZIKV neutralizing antibody titers have expanded clones of memory B cells that express the same immunoglobulin VH3-23/VK1-5 genes. These recurring antibodies cross-react with DENV1, but not other flaviviruses, neutralize both DENV1 and ZIKV, and protect mice against ZIKV challenge. Structural analyses reveal the mechanism of recognition of the ZEDIII lateral ridge by VH3-23/VK1-5 antibodies. Serologic testing shows that antibodies to this region correlate with serum neutralizing activity to ZIKV. Thus, high neutralizing responses to ZIKV are associated with pre-existing reactivity to DENV1 in humans.
- Laboratory of Molecular Immunology, The Rockefeller University, New York, NY 10065, USA. Electronic address: drobbiani@rockefeller.edu.
Organizational Affiliation: