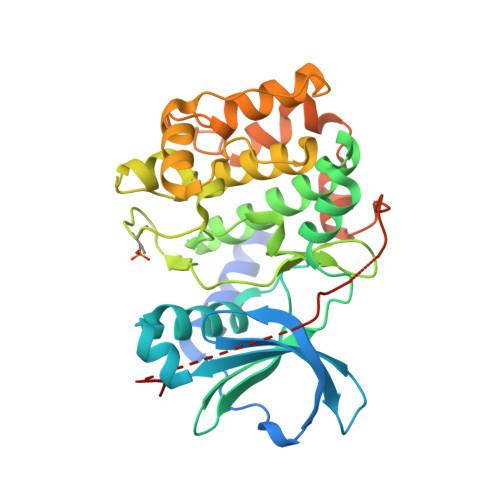
Design of Aminobenzothiazole Inhibitors of Rho Kinases 1 and 2 by Using Protein Kinase A as a Structure Surrogate.
Judge, R.A., Vasudevan, A., Scott, V.E., Simler, G.H., Pratt, S.D., Namovic, M.T., Putman, C.B., Aguirre, A., Stoll, V.S., Mamo, M., Swann, S.I., Cassar, S.C., Faltynek, C.R., Kage, K.L., Boyce-Rustay, J.M., Hobson, A.D.(2018) Chembiochem 19: 613-621
- PubMed: 29314498 Search on PubMed
- DOI: https://doi.org/10.1002/cbic.201700547
- Primary Citation Related Structures:
5VHB, 5VI9, 5VIB - PubMed Abstract:
We describe the design, synthesis, and structure-activity relationships (SARs) of a series of 2-aminobenzothiazole inhibitors of Rho kinases (ROCKs) 1 and 2, which were optimized to low nanomolar potencies by use of protein kinase A (PKA) as a structure surrogate to guide compound design. A subset of these molecules also showed robust activity in a cell-based myosin phosphatase assay and in a mechanical hyperalgesia in vivo pain model.
- AbbVie, Inc., 1 North Waukegan Road, North Chicago, IL, 60064, USA.
Organizational Affiliation: