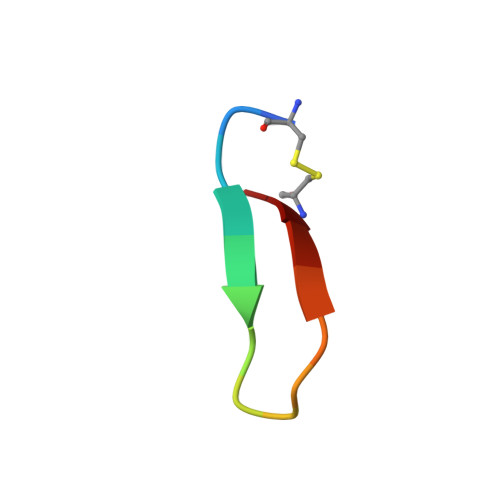
Utilization of peptide phage display to investigate hotspots on IL-17A and what it means for drug discovery.
Ting, J.P., Tung, F., Antonysamy, S., Wasserman, S., Jones, S.B., Zhang, F.F., Espada, A., Broughton, H., Chalmers, M.J., Woodman, M.E., Bina, H.A., Dodge, J.A., Benach, J., Zhang, A., Groshong, C., Manglicmot, D., Russell, M., Afshar, S.(2018) PLoS One 13: e0190850-e0190850
- PubMed: 29329326 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0190850
- Primary Citation Related Structures:
5VB9 - PubMed Abstract:
To date, IL-17A antibodies remain the only therapeutic approach to correct the abnormal activation of the IL-17A/IL-17R signaling complex. Why is it that despite the remarkable success of IL-17 antibodies, there is no small molecule antagonist of IL-17A in the clinic? Here we offer a unique approach to address this question. In order to understand the interaction of IL-17A with its receptor, we combined peptide discovery using phage display with HDX, crystallography, and functional assays to map and characterize hot regions that contribute to most of the energetics of the IL-17A/IL-17R interaction. These functional maps are proposed to serve as a guide to aid in the development of small molecules that bind to IL-17A and block its interaction with IL-17RA.
- Department of protein Engineering, Eli Lilly Biotechnology Center, San Diego, California, United States of America.
Organizational Affiliation: