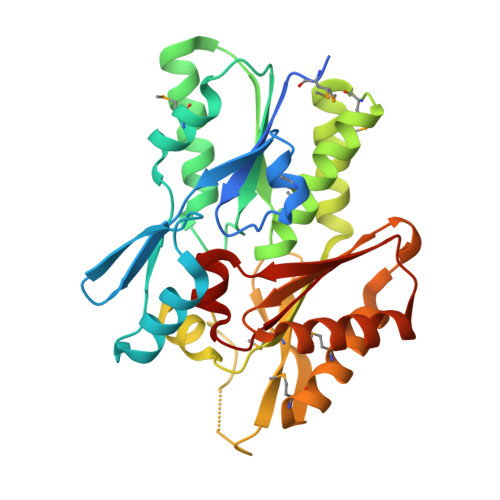
The crystal structure of the Staphylococcus aureus Fatty acid Kinase (Fak) B1 protein loaded with palmitic acid to 1.83 Angstroem resolution
Cuypers, M.G., Ericson, M., Subramanian, C., Broussard, T.C., Miller, D.J., White, S.W., Rock, C.O.(2018) J Biological Chem