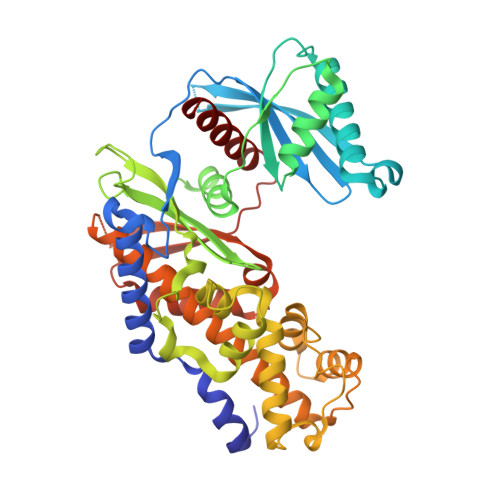
Discovery of potent and orally active 1,4-disubstituted indazoles as novel allosteric glucokinase activators.
Cheruvallath, Z.S., Gwaltney, S.L., Sabat, M., Tang, M., Wang, H., Jennings, A., Hosfield, D., Lee, B., Wu, Y., Halkowycz, P., Grimshaw, C.E.(2017) Bioorg Med Chem Lett 27: 2678-2682
- PubMed: 28512030 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2017.04.041
- Primary Citation Related Structures:
5V4W, 5V4X - PubMed Abstract:
Guided by co-crystal structural information obtained from a different series we were exploring, a scaffold morphing and SBDD approach led to the discovery of the 1,4-disubstituted indazole series as a novel class of GKAs that potently activate GK in enzyme and cell assays. anti-diabetic OGTT efficacy was demonstrated with 29 in a rodent models of type 2 diabetes.
- Takeda California, 10410 Science Center Drive, San Diego 92121, USA. Electronic address: zacharia.cheruvallath@takedasd.com.
Organizational Affiliation: