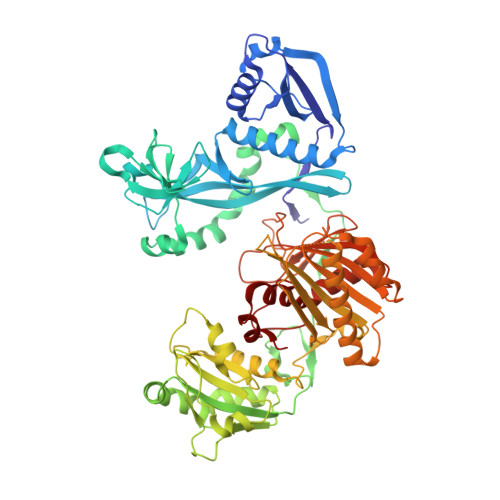
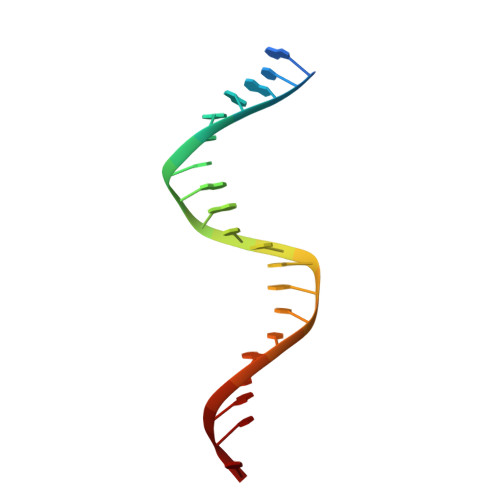
DNA recognition by an RNA-guided bacterial Argonaute.
Doxzen, K.W., Doudna, J.A.(2017) PLoS One 12: e0177097-e0177097
- PubMed: 28520746 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0177097
- Primary Citation Related Structures:
5UX0 - PubMed Abstract:
Argonaute (Ago) proteins are widespread in prokaryotes and eukaryotes and share a four-domain architecture capable of RNA- or DNA-guided nucleic acid recognition. Previous studies identified a prokaryotic Argonaute protein from the eubacterium Marinitoga piezophila (MpAgo), which binds preferentially to 5'-hydroxylated guide RNAs and cleaves single-stranded RNA (ssRNA) and DNA (ssDNA) targets. Here we present a 3.2 Å resolution crystal structure of MpAgo bound to a 21-nucleotide RNA guide and a complementary 21-nucleotide ssDNA substrate. Comparison of this ternary complex to other target-bound Argonaute structures reveals a unique orientation of the N-terminal domain, resulting in a straight helical axis of the entire RNA-DNA heteroduplex through the central cleft of the protein. Additionally, mismatches introduced into the heteroduplex reduce MpAgo cleavage efficiency with a symmetric profile centered around the middle of the helix. This pattern differs from the canonical mismatch tolerance of other Argonautes, which display decreased cleavage efficiency for substrates bearing sequence mismatches to the 5' region of the guide strand. This structural analysis of MpAgo bound to a hybrid helix advances our understanding of the diversity of target recognition mechanisms by Argonaute proteins.
- Biophysics Graduate Group, University of California, Berkeley, California, United States of America.
Organizational Affiliation: