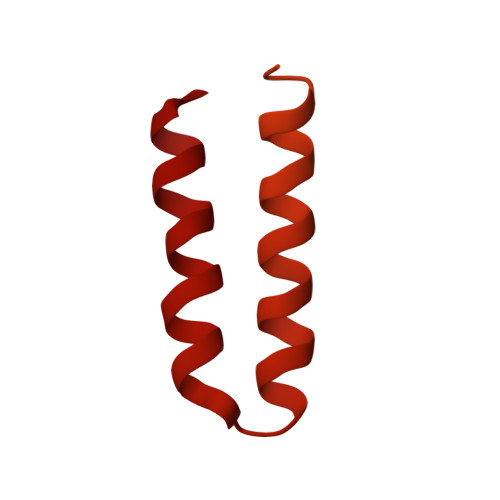
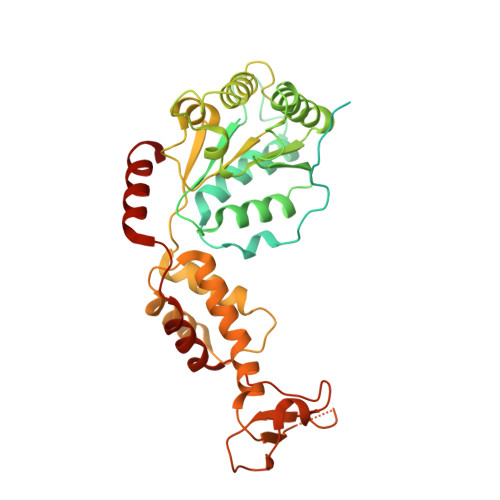
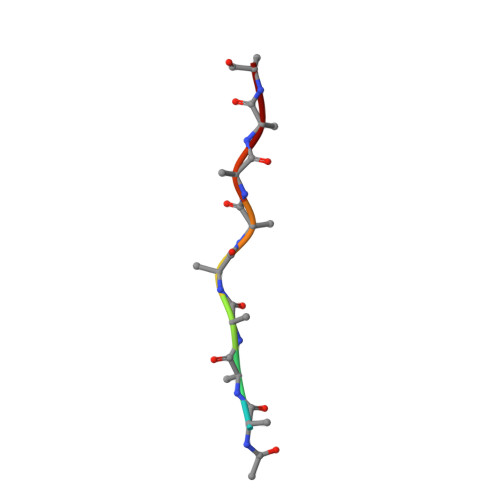
Structural basis of protein translocation by the Vps4-Vta1 AAA ATPase.
Monroe, N., Han, H., Shen, P.S., Sundquist, W.I., Hill, C.P.(2017) Elife 6
- PubMed: 28379137 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.24487
- Primary Citation Related Structures:
5UIE - PubMed Abstract:
Many important cellular membrane fission reactions are driven by ESCRT pathways, which culminate in disassembly of ESCRT-III polymers by the AAA ATPase Vps4. We report a 4.3 Å resolution cryo-EM structure of the active Vps4 hexamer with its cofactor Vta1, ADP·BeF x , and an ESCRT-III substrate peptide. Four Vps4 subunits form a helix whose interfaces are consistent with ATP binding, is stabilized by Vta1, and binds the substrate peptide. The fifth subunit approximately continues this helix but appears to be dissociating. The final Vps4 subunit completes a notched-washer configuration as if transitioning between the ends of the helix. We propose that ATP binding propagates growth at one end of the helix while hydrolysis promotes disassembly at the other end, so that Vps4 'walks' along ESCRT-III until it encounters the ordered N-terminal domain to destabilize the ESCRT-III lattice. This model may be generally applicable to other protein-translocating AAA ATPases.
- Department of Biochemistry, University of Utah School of Medicine, Salt Lake City, United States.
Organizational Affiliation: