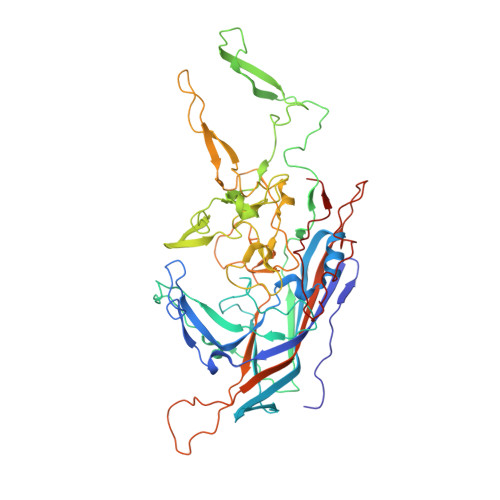
The 2.8 angstrom Electron Microscopy Structure of Adeno-Associated Virus-DJ Bound by a Heparinoid Pentasaccharide.
Xie, Q., Spear, J.M., Noble, A.J., Sousa, D.R., Meyer, N.L., Davulcu, O., Zhang, F., Linhardt, R.J., Stagg, S.M., Chapman, M.S.(2017) Mol Ther Methods Clin Dev 5: 1-12
- PubMed: 28480299 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.omtm.2017.02.004
- Primary Citation Related Structures:
5UF6 - PubMed Abstract:
Atomic structures of adeno-associated virus (AAV)-DJ, alone and in complex with fondaparinux, have been determined by cryoelectron microscopy at 3 Å resolution. The gene therapy vector, AAV-DJ, is a hybrid of natural serotypes that was previously derived by directed evolution, selecting for hepatocyte entry and resistance to neutralization by human serum. The structure of AAV-DJ differs from that of parental serotypes in two regions where neutralizing antibodies bind, so immune escape appears to have been the primary driver of AAV-DJ's directed evolution. Fondaparinux is an analog of cell surface heparan sulfate to which several AAVs bind during entry. Fondaparinux interacts with viral arginines at a known heparin binding site, without the large conformational changes whose presence was controversial in low-resolution imaging of AAV2-heparin complexes. The glycan density suggests multi-modal binding that could accommodate sequence variation and multivalent binding along a glycan polymer, consistent with a role in attachment, prior to more specific interactions with a receptor protein mediating entry.
- Department of Biochemistry & Molecular Biology, School of Medicine, Oregon Health & Science University, Portland, OR 97239-3098, USA.
Organizational Affiliation: