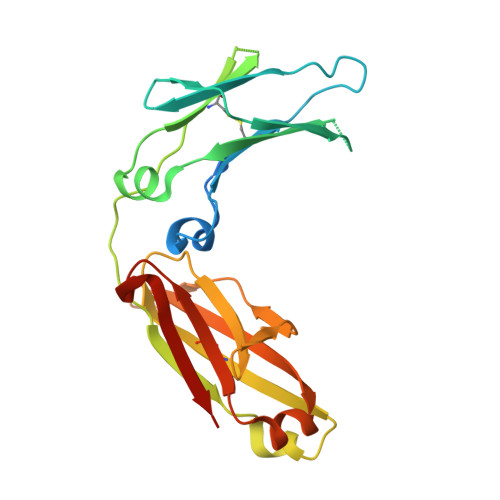
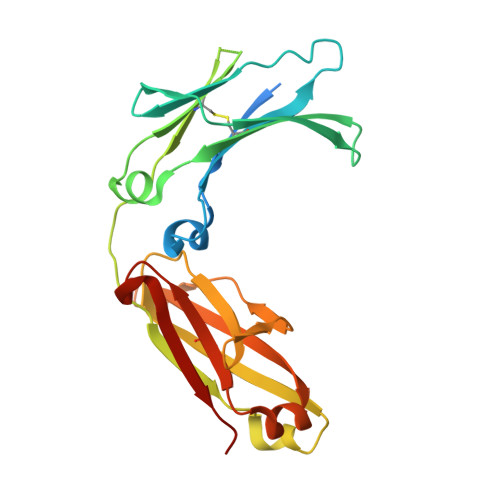
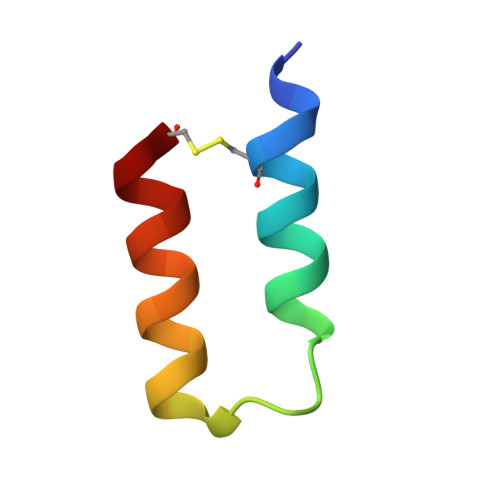
Modulation of protein A binding allows single-step purification of mouse bispecific antibodies that retain FcRn binding.
Zwolak, A., Armstrong, A.A., Tam, S.H., Pardinas, J.R., Goulet, D.R., Zheng, S., Brosnan, K., Emmell, E., Luo, J., Gilliland, G.L., Chiu, M.L.(2017) MAbs 9: 1306-1316
- PubMed: 28898162 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1080/19420862.2017.1375639
- Primary Citation Related Structures:
5UBX - PubMed Abstract:
The increased number of bispecific antibodies (BsAb) under therapeutic development has resulted in a need for mouse surrogate BsAbs. Here, we describe a one-step method for generating highly pure mouse BsAbs suitable for in vitro and in vivo studies. We identify two mutations in the mouse IgG2a and IgG2b Fc region: one that eliminates protein A binding and one that enhances protein A binding by 8-fold. We show that BsAbs harboring these mutations can be purified from the residual parental monoclonal antibodies in one step using protein A affinity chromatography. The structural basis for the effects of these mutations was analyzed by X-ray crystallography. While the mutation that disrupted protein A binding also inhibited FcRn interaction, a bispecific mutant in which one subunit retained the ability to bind protein A could still interact with FcRn. Pharmacokinetic analysis of the serum half-lives of the mutants showed that the mutant BsAb had a serum half-life comparable to a wild-type Ab. The results describe a rapid method for generating panels of mouse BsAbs that could be used in mouse studies.
- a Biologics Discovery , Janssen Research & Development, LLC , Spring House , PA , USA.
Organizational Affiliation: