UbcH5a-Ub isopeptide conjugate
Sanchez, J.G., Wagner, J.M., Pornillos, O.To be published.
Experimental Data Snapshot
Starting Models: experimental
View more details
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Ubiquitin-conjugating enzyme E2 D1 | 150 | Homo sapiens | Mutation(s): 1 Gene Names: UBE2D1, SFT, UBC5A, UBCH5, UBCH5A EC: 2.3.2.23 (PDB Primary Data), 2.3.2.24 (PDB Primary Data) | 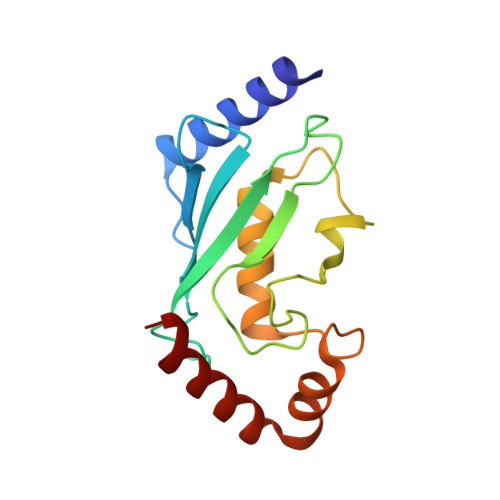 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P51668 GTEx: ENSG00000072401 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P51668 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Ubiquitin | 76 | Homo sapiens | Mutation(s): 0 Gene Names: UBB | 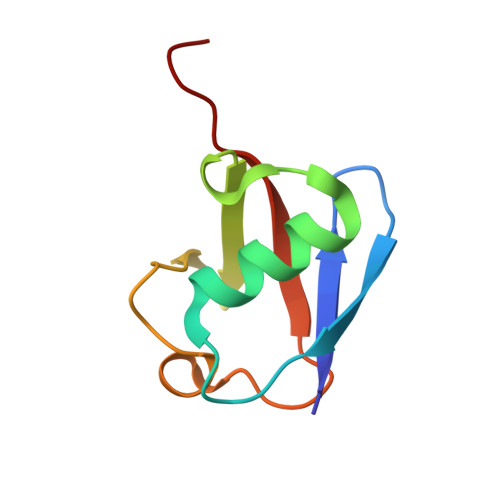 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P0CG47 GTEx: ENSG00000170315 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P0CG47 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 41.949 | α = 90 |
| b = 51.41 | β = 101.39 |
| c = 53.157 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| HKL-2000 | data reduction |
| HKL-2000 | data scaling |
| PHENIX | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Institutes of Health/National Institute of General Medical Sciences (NIH/NIGMS) | United States | R01-GM112508 |