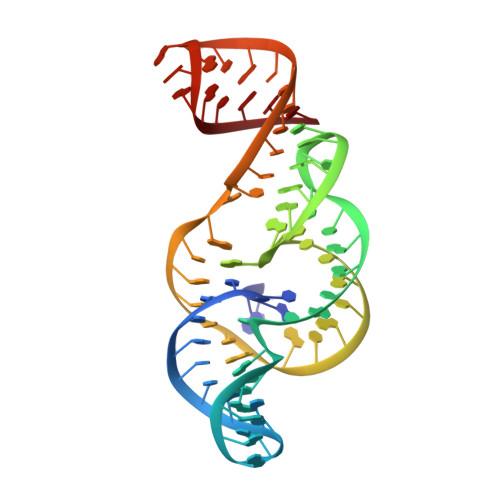
Zika virus produces noncoding RNAs using a multi-pseudoknot structure that confounds a cellular exonuclease.
Akiyama, B.M., Laurence, H.M., Massey, A.R., Costantino, D.A., Xie, X., Yang, Y., Shi, P.Y., Nix, J.C., Beckham, J.D., Kieft, J.S.(2016) Science 354: 1148-1152
- PubMed: 27934765 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.aah3963
- Primary Citation Related Structures:
5TPY - PubMed Abstract:
The outbreak of Zika virus (ZIKV) and associated fetal microcephaly mandates efforts to understand the molecular processes of infection. Related flaviviruses produce noncoding subgenomic flaviviral RNAs (sfRNAs) that are linked to pathogenicity in fetal mice. These viruses make sfRNAs by co-opting a cellular exonuclease via structured RNAs called xrRNAs. We found that ZIKV-infected monkey and human epithelial cells, mouse neurons, and mosquito cells produce sfRNAs. The RNA structure that is responsible for ZIKV sfRNA production forms a complex fold that is likely found in many pathogenic flaviviruses. Mutations that disrupt the structure affect exonuclease resistance in vitro and sfRNA formation during infection. The complete ZIKV xrRNA structure clarifies the mechanism of exonuclease resistance and identifies features that may modulate function in diverse flaviviruses.
- Department of Biochemistry and Molecular Genetics, University of Colorado Denver School of Medicine, Aurora, CO 80045, USA.
Organizational Affiliation: