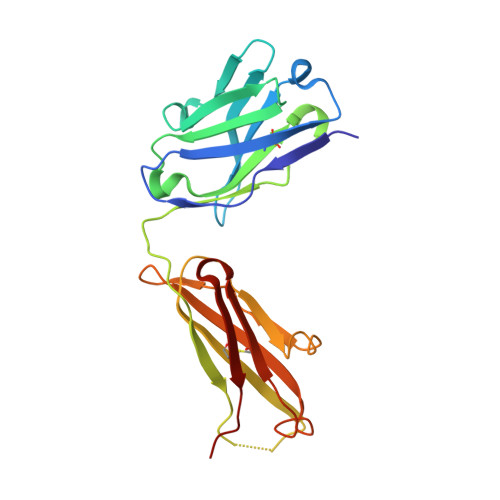
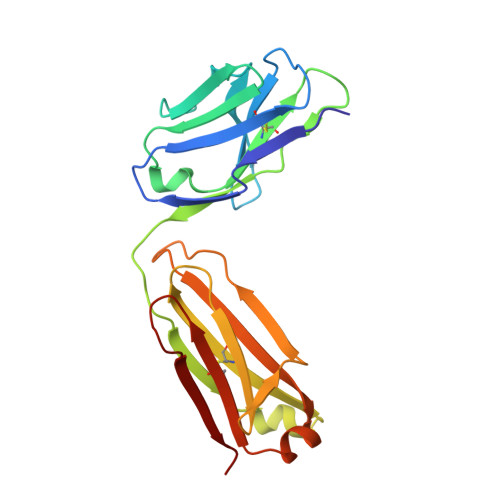
Structural basis for inhibition of erythrocyte invasion by antibodies toPlasmodium falciparumprotein CyRPA.
Chen, L., Xu, Y., Wong, W., Thompson, J.K., Healer, J., Goddard-Borger, E.D., Lawrence, M.C., Cowman, A.F.(2017) Elife 6
- PubMed: 28195530 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.21347
- Primary Citation Related Structures:
5TIH, 5TIK - PubMed Abstract:
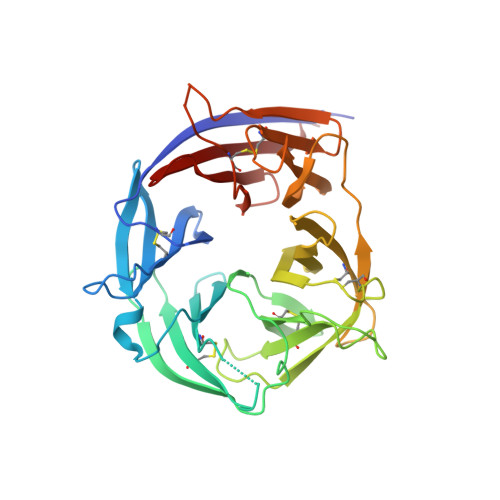
Plasmodium falciparum causes malaria in humans with over 450,000 deaths annually. The asexual blood stage involves invasion of erythrocytes by merozoites, in which they grow and divide to release daughter merozoites, which in turn invade new erythrocytes perpetuating the cycle responsible for malaria. A key step in merozoite invasion is the essential binding of PfRh5/CyRPA/PfRipr complex to basigin, a step linked to the formation of a pore between merozoites and erythrocytes. We show CyRPA interacts directly with PfRh5. An invasion inhibitory monoclonal antibody to CyRPA blocks binding of CyRPA to PfRh5 and complex formation thus illuminating the molecular mechanism for inhibition of parasite growth. We determined the crystal structures of CyRPA alone and in complex with an antibody Fab fragment. CyRPA has a six-bladed β-propeller fold, and we identify the region that interacts with PfRh5. This functionally conserved epitope is a potential target for vaccines against P. falciparum .
- The Walter and Eliza Hall Institute of Medical Research, Melbourne, Australia.
Organizational Affiliation: