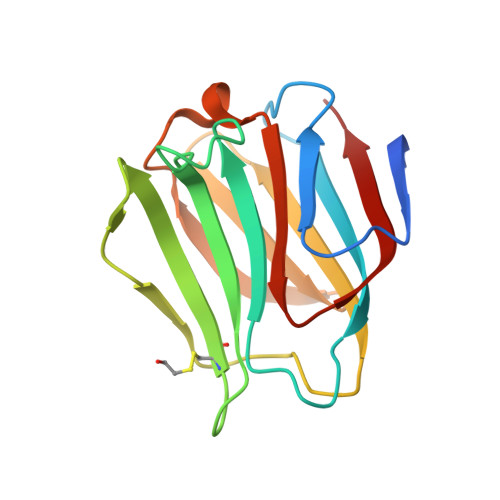
Structure-based rationale for differential recognition of lacto- and neolacto- series glycosphingolipids by the N-terminal domain of human galectin-8.
Bohari, M.H., Yu, X., Zick, Y., Blanchard, H.(2016) Sci Rep 6: 39556-39556
- PubMed: 28000747 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep39556
- Primary Citation Related Structures:
5T7I, 5T7S, 5T7T, 5T7U - PubMed Abstract:
Glycosphingolipids are ubiquitous cell surface molecules undertaking fundamental cellular processes. Lacto-N-tetraose (LNT) and lacto-N-neotetraose (LNnT) are the representative core structures for lacto- and neolacto-series glycosphingolipids. These glycolipids are the carriers to the blood group antigen and human natural killer antigens mainly found on blood cells, and are also principal components in human milk, contributing to infant health. The β-galactoside recognising galectins mediate various cellular functions of these glycosphingolipids. We report crystallographic structures of the galectin-8 N-terminal domain (galectin-8N) in complex with LNT and LNnT. We reveal the first example in which the non-reducing end of LNT binds to the primary binding site of a galectin, and provide a structure-based rationale for the significant ten-fold difference in binding affinities of galectin-8N toward LNT compared to LNnT, such a magnitude of difference not being observed for any other galectin. In addition, the LNnT complex showed that the unique Arg59 has ability to adopt a new orientation, and comparison of glycerol- and lactose-bound galectin-8N structures reveals a minimum atomic framework for ligand recognition. Overall, these results enhance our understanding of glycosphingolipids interactions with galectin-8N, and highlight a structure-based rationale for its significantly different affinity for components of biologically relevant glycosphingolipids.
- Institute for Glycomics, Griffith University, Gold Coast Campus, 4222, Australia.
Organizational Affiliation: