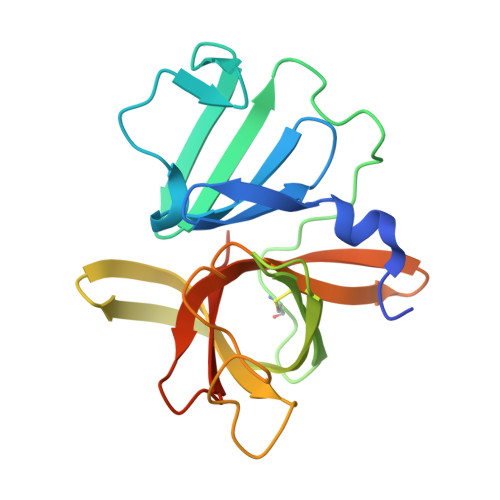
Structure-based exploration and exploitation of the S4 subsite of norovirus 3CL protease in the design of potent and permeable inhibitors.
Galasiti Kankanamalage, A.C., Kim, Y., Rathnayake, A.D., Damalanka, V.C., Weerawarna, P.M., Doyle, S.T., Alsoudi, A.F., Dissanayake, D.M., Lushington, G.H., Mehzabeen, N., Battaile, K.P., Lovell, S., Chang, K.O., Groutas, W.C.(2016) Eur J Med Chem 126: 502-516
- PubMed: 27914364 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.ejmech.2016.11.027
- Primary Citation Related Structures:
5T6D, 5T6F, 5T6G - PubMed Abstract:
Human noroviruses are the primary cause of epidemic and sporadic acute gastroenteritis. The worldwide high morbidity and mortality associated with norovirus infections, particularly among the elderly, immunocompromised patients and children, constitute a serious public health concern. There are currently no approved human vaccines or norovirus-specific small-molecule therapeutics or prophylactics. Norovirus 3CL protease has recently emerged as a potential therapeutic target for the development of anti-norovirus agents. We hypothesized that the S 4 subsite of the enzyme may provide an effective means of designing potent and cell permeable inhibitors of the enzyme. We report herein the structure-guided exploration and exploitation of the S 4 subsite of norovirus 3CL protease in the design and synthesis of effective inhibitors of the protease.
- Department of Chemistry, Wichita State University, Wichita, KS 67260, USA.
Organizational Affiliation: