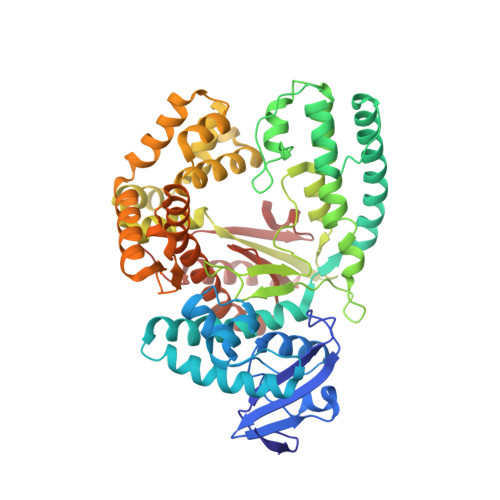
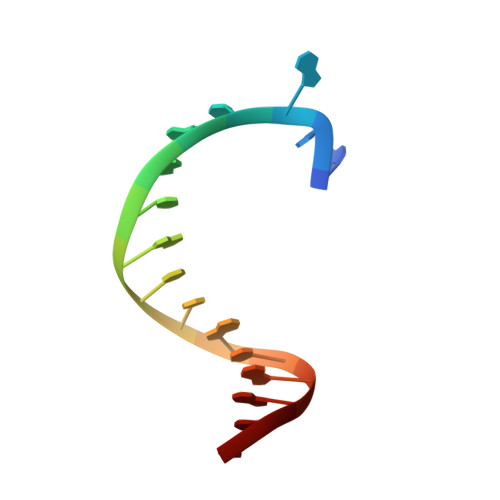
Structural Basis for the KlenTaq DNA Polymerase Catalysed Incorporation of Alkene- versus Alkyne-Modified Nucleotides.
Hottin, A., Betz, K., Diederichs, K., Marx, A.(2017) Chemistry 23: 2109-2118
- PubMed: 27901305 Search on PubMed
- DOI: https://doi.org/10.1002/chem.201604515
- Primary Citation Related Structures:
5E41, 5SZT - PubMed Abstract:
Efficient incorporation of modified nucleotides by DNA polymerases is essential for many cutting-edge biomolecular technologies. The present study compares the acceptance of either alkene- or alkyne-modified nucleotides by KlenTaq DNA polymerase and provides structural insights into how 7-deaza-adenosine and deoxyuridine with attached alkene-modifications are incorporated into the growing DNA strand. Thereby, we identified modified nucleotides that prove to be superior substrates for KlenTaq DNA polymerase compared with their natural analogues. The knowledge can be used to guide future design of functionalized nucleotide building blocks.
- Department of Chemistry and Department of Biology, University of Konstanz, Universitätsstrasse 10, 78457, Konstanz, Germany.
Organizational Affiliation: