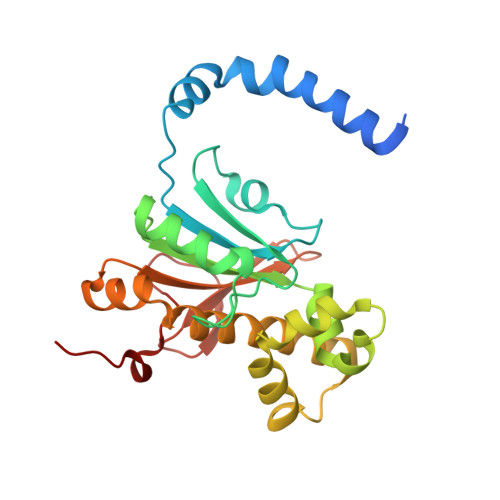
The structure, kinetics and interactions of the beta-carboxysomal beta-carbonic anhydrase, CcaA.
McGurn, L.D., Moazami-Goudarzi, M., White, S.A., Suwal, T., Brar, B., Tang, J.Q., Espie, G.S., Kimber, M.S.(2016) Biochem J 473: 4559-4572
- PubMed: 27729545 Search on PubMed
- DOI: https://doi.org/10.1042/BCJ20160773
- Primary Citation Related Structures:
5SWC - PubMed Abstract:
CcaA is a β-carbonic anhydrase (CA) that is a component of the carboxysomes of a subset of β-cyanobacteria. This protein, which has a characteristic C-terminal extension of unknown function, is recruited to the carboxysome via interactions with CcmM, which is itself a γ-CA homolog with enzymatic activity in many, but not all cyanobacteria. We have determined the structure of CcaA from Synechocystis sp. PCC 6803 at 1.45 Å. In contrast with the dimer-of-dimers organization of most bacterial β-CAs, or the loose dimer-of-dimers-of-dimers organization found in the plant enzymes, CcaA shows a well-packed trimer-of-dimers organization. The proximal part of the characteristic C-terminal extension is ordered by binding at a site that passes through the two-fold symmetry axis shared with an adjacent dimer; as a result, only one of a pair of converging termini can be ordered at any given time. Docking in Rosetta failed to find well-packed solutions, indicating that formation of the CcaA/CcmM complex probably requires significant backbone movements in at least one of the binding partners. Surface plasmon resonance experiments showed that CcaA forms a complex with CcmM with sub-picomolar affinity, with contributions from residues in CcmM's αA helix and CcaA's C-terminal tail. Catalytic characterization showed CcaA to be among the least active β-CAs characterized to date, with activity comparable with the γ-CA, CcmM, it either complements or replaces. Intriguingly, the C-terminal tail appears to partly inhibit activity, possibly indicating a role in minimizing the activity of unencapsulated enzyme.
- Department of Molecular and Cellular Biology, University of Guelph, Guelph, Ontario, Canada N1G2W1.
Organizational Affiliation: