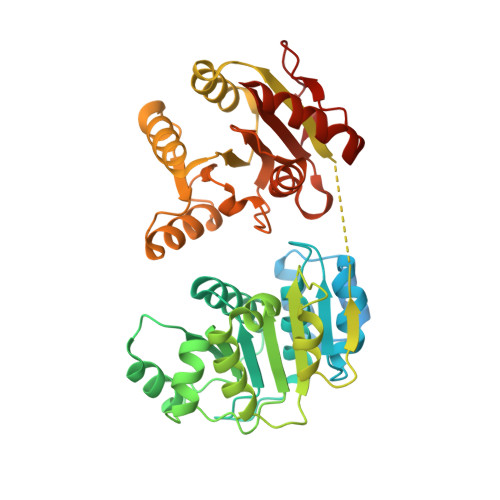
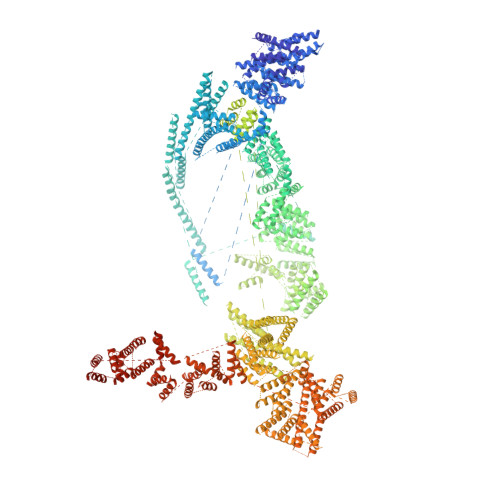
Structural and biochemical analyses of the DEAD-box ATPase Sub2 in association with THO or Yra1.
Ren, Y., Schmiege, P., Blobel, G.(2017) Elife 6
- PubMed: 28059701 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.20070
- Primary Citation Related Structures:
5SUP, 5SUQ - PubMed Abstract:
mRNA is cotranscrptionally processed and packaged into messenger ribonucleoprotein particles (mRNPs) in the nucleus. Prior to export through the nuclear pore, mRNPs undergo several obligatory remodeling reactions. In yeast, one of these reactions involves loading of the mRNA-binding protein Yra1 by the DEAD-box ATPase Sub2 as assisted by the hetero-pentameric THO complex. To obtain molecular insights into reaction mechanisms, we determined crystal structures of two relevant complexes: a THO hetero-pentamer bound to Sub2 at 6.0 Å resolution; and Sub2 associated with an ATP analogue, RNA, and a C-terminal fragment of Yra1 (Yra1-C) at 2.6 Å resolution. We found that the 25 nm long THO clamps Sub2 in a half-open configuration; in contrast, when bound to the ATP analogue, RNA and Yra1-C, Sub2 assumes a closed conformation. Both THO and Yra1-C stimulated Sub2's intrinsic ATPase activity. We propose that THO surveys common landmarks in each nuclear mRNP to localize Sub2 for targeted loading of Yra1.
- Laboratory of Cell Biology, Howard Hughes Medical Institute, The Rockefeller University, New York, United States.
Organizational Affiliation: