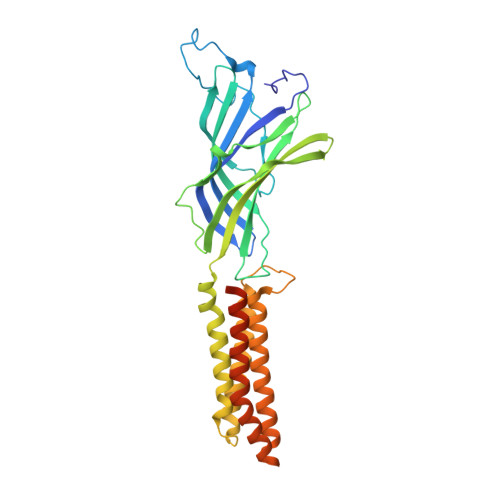
Crystal structures of a GABAA-receptor chimera reveal new endogenous neurosteroid-binding sites.
Laverty, D., Thomas, P., Field, M., Andersen, O.J., Gold, M.G., Biggin, P.C., Gielen, M., Smart, T.G.(2017) Nat Struct Mol Biol 24: 977-985
- PubMed: 28967882 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.3477
- Primary Citation Related Structures:
5OSA, 5OSB, 5OSC - PubMed Abstract:
γ-Aminobutyric acid receptors (GABA A Rs) are vital for controlling excitability in the brain. This is emphasized by the numerous neuropsychiatric disorders that result from receptor dysfunction. A critical component of most native GABA A Rs is the α subunit. Its transmembrane domain is the target for many modulators, including endogenous brain neurosteroids that impact anxiety, stress and depression, and for therapeutic drugs, such as general anesthetics. Understanding the basis for the modulation of GABA A R function requires high-resolution structures. Here we present the first atomic structures of a GABA A R chimera at 2.8-Å resolution, including those bound with potentiating and inhibitory neurosteroids. These structures define new allosteric binding sites for these modulators that are associated with the α-subunit transmembrane domain. Our findings will enable the exploitation of neurosteroids for therapeutic drug design to regulate GABA A Rs in neurological disorders.
- Department of Neuroscience, Physiology & Pharmacology, University College London, London, UK.
Organizational Affiliation: