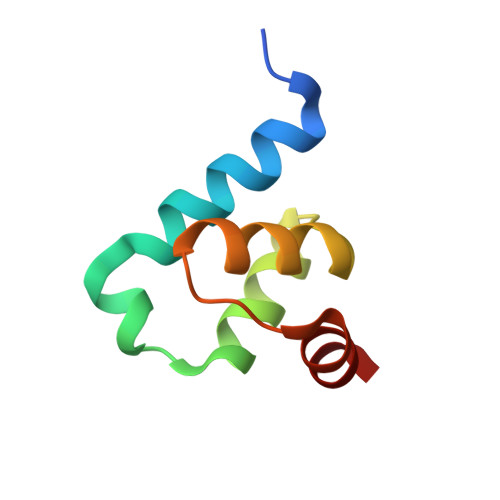
Crystal structure of Trypanosoma Brucei PEX14 N-terminal domain in complex with small molecules to investigate the water envelope
Ratkova, E.L., Dawidowski, M., Napolitano, V., Dubin, G., Fino, R., Popowicz, G., Sattler, M., Tetko, I.V.To be published.