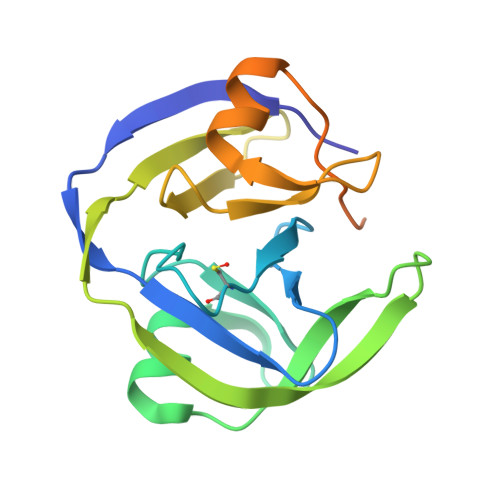
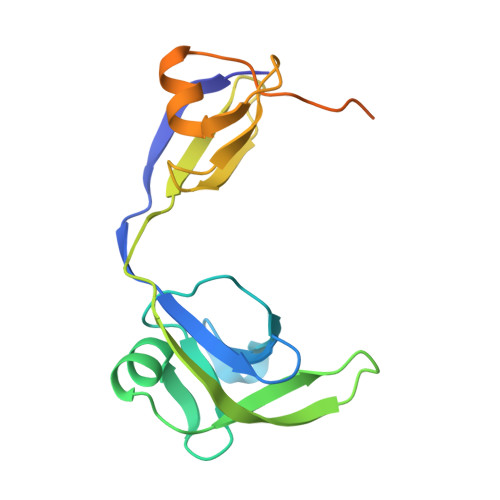
Mechanistic Insights into Cyclic Peptide Generation by DnaE Split-Inteins through Quantitative and Structural Investigation.
Kick, L.M., Harteis, S., Koch, M.F., Schneider, S.(2017) Chembiochem 18: 2242-2246
- PubMed: 28914478 Search on PubMed
- DOI: https://doi.org/10.1002/cbic.201700503
- Primary Citation Related Structures:
5OL1, 5OL5, 5OL6, 5OL7 - PubMed Abstract:
Inteins carry out protein-splicing reactions, which are used in protein chemistry, protein engineering and biotechnological applications. Rearrangement of the order of the domains in split-inteins results in head-to-tail cyclisation of the target sequence, which can be used for genetic encoding and expression of libraries of cyclic peptides (CPs). The efficiency of the splicing reaction depends on the target sequence. Here we used mass spectrometry to assess in vivo cyclic peptide formation from different hexameric target sequences by the DnaE split-inteins from Synechocystis sp. and Nostoc punctiforme, revealing a strong impact of the target sequence and of the intein on the intracellular peptide concentration. Furthermore, we determined the crystal structures of their pre-splicing complexes, which allowed us to identify F-block Asp17 as crucial for the DnaE-mediated splicing reaction.
- Center for Integrated Protein Science, Department of Chemistry, Technische Universität München, Lichtenbergstrasse 4, 85748, Garching, Germany.
Organizational Affiliation: