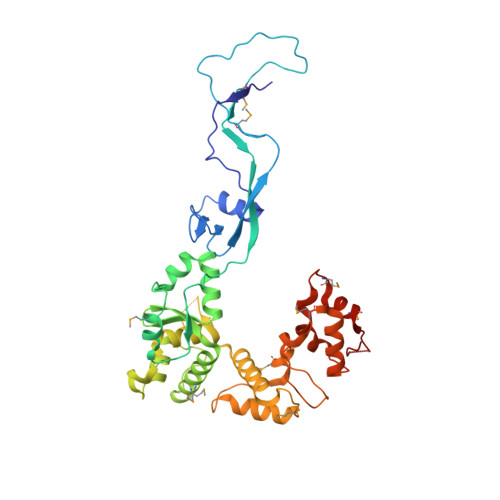
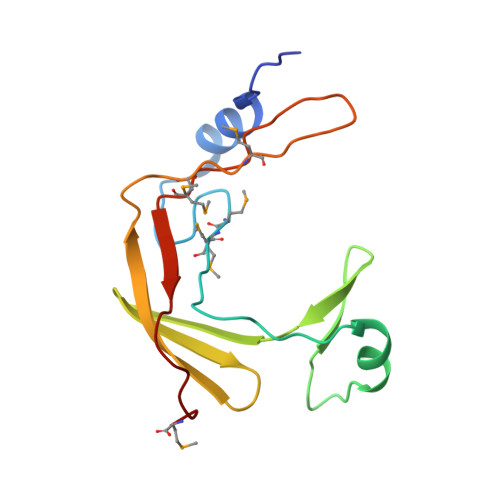
Structural Basis for the Recruitment of Ctf18-RFC to the Replisome.
Grabarczyk, D.B., Silkenat, S., Kisker, C.(2018) Structure 26: 137-144.e3
- PubMed: 29225079 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2017.11.004
- Primary Citation Related Structures:
5OKC, 5OKI - PubMed Abstract:
Ctf18-RFC is an alternative PCNA loader which plays important but poorly understood roles in multiple DNA replication-associated processes. To fulfill its specialist roles, the Ctf18-RFC clamp loader contains a unique module in which the Dcc1-Ctf8 complex is bound to the C terminus of Ctf18 (the Ctf18-1-8 module). Here, we report the structural and functional characterization of the heterotetrameric complex formed between Ctf18-1-8 and a 63 kDa fragment of DNA polymerase ɛ. Our data reveal that Ctf18-1-8 binds stably to the polymerase and far from its other functional sites, suggesting that Ctf18-RFC could be associated with Pol ɛ throughout normal replication as the leading strand clamp loader. We also show that Pol ɛ and double-stranded DNA compete to bind the same winged-helix domain on Dcc1, with Pol ɛ being the preferred binding partner, thus suggesting that there are two alternative pathways to recruit Ctf18-RFC to sites of replication.
- Rudolf Virchow Center for Experimental Biomedicine, Institute for Structural Biology, Josef-Schneider-Strasse 2, 97080 Würzburg, Germany. Electronic address: daniel.grabarczyk@virchow.uni-wuerzburg.de.
Organizational Affiliation: