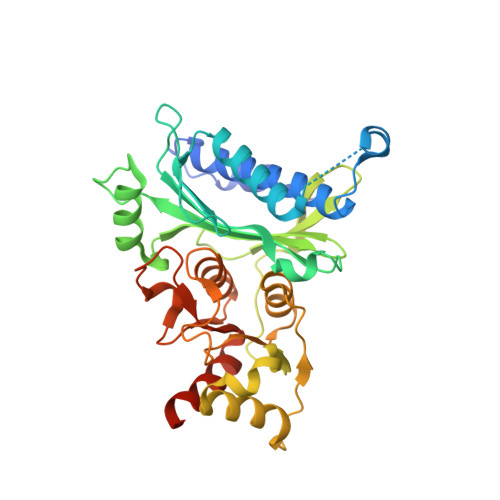
Structures of Leishmania Fructose-1,6-Bisphosphatase Reveal Species-Specific Differences in the Mechanism of Allosteric Inhibition.
Yuan, M., Vasquez-Valdivieso, M.G., McNae, I.W., Michels, P.A.M., Fothergill-Gilmore, L.A., Walkinshaw, M.D.(2017) J Mol Biology 429: 3075-3089
- PubMed: 28882541 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2017.08.010
- Primary Citation Related Structures:
5OEY, 5OEZ, 5OFU - PubMed Abstract:
The gluconeogenic enzyme fructose-1,6-bisphosphatase has been proposed as a potential drug target against Leishmania parasites that cause up to 20,000-30,000 deaths annually. A comparison of three crystal structures of Leishmania major fructose-1,6-bisphosphatase (LmFBPase) along with enzyme kinetic data show how AMP acts as an allosteric inhibitor and provides insight into its metal-dependent reaction mechanism. The crystal structure of the apoenzyme form of LmFBPase is a homotetramer in which the dimer of dimers adopts a planar conformation with disordered "dynamic loops". The structure of LmFBPase, complexed with manganese and its catalytic product phosphate, shows the dynamic loops locked into the active sites. A third crystal structure of LmFBPase complexed with its allosteric inhibitor AMP shows an inactive form of the tetramer, in which the dimer pairs are rotated by 18° relative to each other. The three structures suggest an allosteric mechanism in which AMP binding triggers a rearrangement of hydrogen bonds across the large and small interfaces. Retraction of the "effector loop" required for AMP binding releases the side chain of His23 from the dimer-dimer interface. This is coupled with a flip of the side chain of Arg48 which ties down the key catalytic dynamic loop in a disengaged conformation and also locks the tetramer in an inactive rotated T-state. The structure of the effector site of LmFBPase shows different structural features compared with human FBPases, thereby offering a potential and species-specific drug target.
- Centre for Translational and Chemical Biology, School of Biological Sciences, University of Edinburgh, Michael Swann Building, Max Born Crescent, Edinburgh EH9 3BF, UK.
Organizational Affiliation: