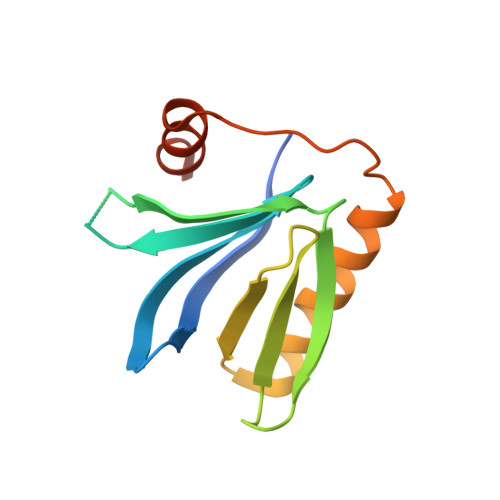
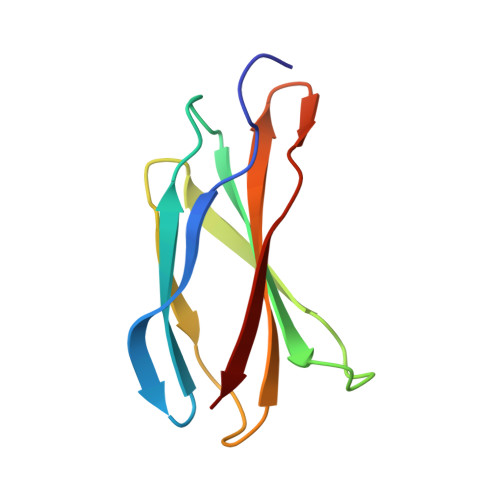
Structural and functional dissection of the DH and PH domains of oncogenic Bcr-Abl tyrosine kinase.
Reckel, S., Gehin, C., Tardivon, D., Georgeon, S., Kukenshoner, T., Lohr, F., Koide, A., Buchner, L., Panjkovich, A., Reynaud, A., Pinho, S., Gerig, B., Svergun, D., Pojer, F., Guntert, P., Dotsch, V., Koide, S., Gavin, A.C., Hantschel, O.(2017) Nat Commun 8: 2101-2101
- PubMed: 29235475 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-017-02313-6
- Primary Citation Related Structures:
5N6R, 5N7E, 5OC7 - PubMed Abstract:
The two isoforms of the Bcr-Abl tyrosine kinase, p210 and p190, are associated with different leukemias and have a dramatically different signaling network, despite similar kinase activity. To provide a molecular rationale for these observations, we study the Dbl-homology (DH) and Pleckstrin-homology (PH) domains of Bcr-Abl p210, which constitute the only structural differences to p190. Here we report high-resolution structures of the DH and PH domains and characterize conformations of the DH-PH unit in solution. Our structural and functional analyses show no evidence that the DH domain acts as a guanine nucleotide exchange factor, whereas the PH domain binds to various phosphatidylinositol-phosphates. PH-domain mutants alter subcellular localization and result in decreased interactions with p210-selective interaction partners. Hence, the PH domain, but not the DH domain, plays an important role in the formation of the differential p210 and p190 Bcr-Abl signaling networks.
- Swiss Institute for Experimental Cancer Research (ISREC), School of Life Sciences, École polytechnique fédérale de Lausanne (EPFL), 1015, Lausanne, Switzerland.
Organizational Affiliation: