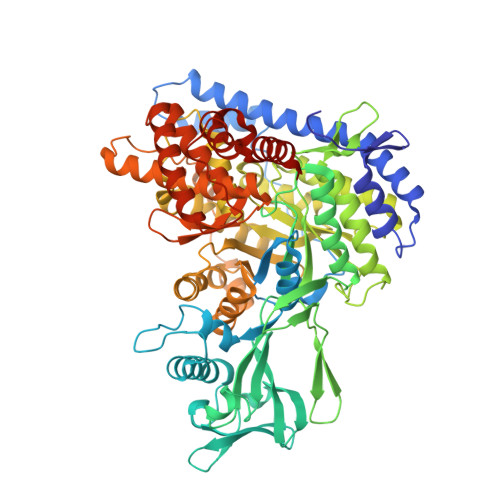
Structural and Functional Characterization of Malate Synthase G from Opportunistic Pathogen Pseudomonas aeruginosa.
McVey, A.C., Medarametla, P., Chee, X., Bartlett, S., Poso, A., Spring, D.R., Rahman, T., Welch, M.(2017) Biochemistry 56: 5539-5549
- PubMed: 28985053 Search on PubMed
- DOI: https://doi.org/10.1021/acs.biochem.7b00852
- Primary Citation Related Structures:
5OAS - PubMed Abstract:
Pseudomonas aeruginosa is an opportunistic human pathogen recognized as a critical threat by the World Health Organization because of the dwindling number of effective therapies available to treat infections. Over the past decade, it has become apparent that the glyoxylate shunt plays a vital role in sustaining P. aeruginosa during infection scenarios. The glyoxylate shunt comprises two enzymes: isocitrate lyase and malate synthase isoform G. Inactivation of these enzymes has been reported to abolish the ability of P. aeruginosa to establish infection in a mammalian model system, yet we still lack the structural information to support drug design efforts. In this work, we describe the first X-ray crystal structure of P. aeruginosa malate synthase G in the apo form at 1.62 Å resolution. The enzyme is a monomer composed of four domains and is highly conserved with homologues found in other clinically relevant microorganisms. It is also dependent on Mg 2+ for catalysis. Metal ion binding led to a change in the intrinsic fluorescence of the protein, allowing us to quantitate its affinity for Mg 2+ . We also identified putative drug binding sites in malate synthase G using computational analysis and, because of the high resolution of the experimental data, were further able to characterize its hydration properties. Our data reveal two promising binding pockets in malate synthase G that may be exploited for drug design.
- Department of Biochemistry, University of Cambridge , Cambridge CB2 1QW, U.K.
Organizational Affiliation: