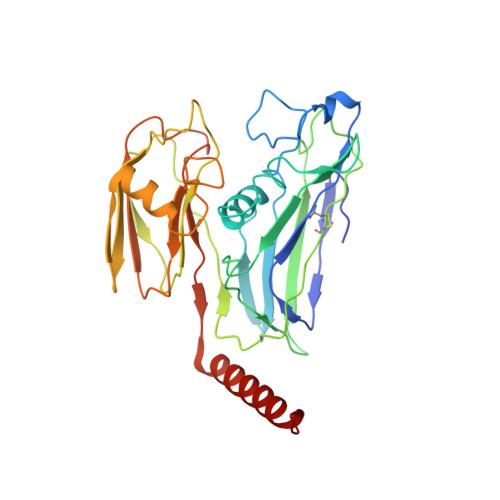
Virus found in a boreal lake links ssDNA and dsDNA viruses.
Laanto, E., Mantynen, S., De Colibus, L., Marjakangas, J., Gillum, A., Stuart, D.I., Ravantti, J.J., Huiskonen, J.T., Sundberg, L.R.(2017) Proc Natl Acad Sci U S A 114: 8378-8383
- PubMed: 28716906 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1703834114
- Primary Citation Related Structures:
5OAC - PubMed Abstract:
Viruses have impacted the biosphere in numerous ways since the dawn of life. However, the evolution, genetic, structural, and taxonomic diversity of viruses remain poorly understood, in part because sparse sampling of the virosphere has concentrated mostly on exploring the abundance and diversity of dsDNA viruses. Furthermore, viral genomes are highly diverse, and using only the current sequence-based methods for classifying viruses and studying their phylogeny is complicated. Here we describe a virus, FLiP ( Flavobacterium -infecting, lipid-containing phage), with a circular ssDNA genome and an internal lipid membrane enclosed in the icosahedral capsid. The 9,174-nt-long genome showed limited sequence similarity to other known viruses. The genetic data imply that this virus might use replication mechanisms similar to those found in other ssDNA replicons. However, the structure of the viral major capsid protein, elucidated at near-atomic resolution using cryo-electron microscopy, is strikingly similar to that observed in dsDNA viruses of the PRD1-adenovirus lineage, characterized by a major capsid protein bearing two β-barrels. The strong similarity between FLiP and another member of the structural lineage, bacteriophage PM2, extends to the capsid organization (pseudo T = 21 dextro ) despite the difference in the genetic material packaged and the lack of significant sequence similarity.
- Centre of Excellence in Biological Interactions, Department of Biological and Environmental Science, University of Jyväskylä, 40014 Jyväskylä, Finland.
Organizational Affiliation: