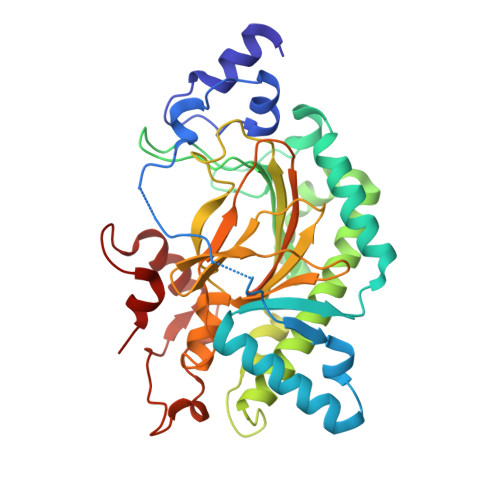
Crystal structure of thebaine 6-O-demethylase from the morphine biosynthesis pathway.
Kluza, A., Niedzialkowska, E., Kurpiewska, K., Wojdyla, Z., Quesne, M., Kot, E., Porebski, P.J., Borowski, T.(2018) J Struct Biol 202: 229-235
- PubMed: 29408320 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2018.01.007
- Primary Citation Related Structures:
5O7Y, 5O9W - PubMed Abstract:
Thebaine 6-O-demethylase (T6ODM) from Papaver somniferum (opium poppy), which belongs to the non-heme 2-oxoglutarate/Fe(II)-dependent dioxygenases (ODD) family, is a key enzyme in the morphine biosynthesis pathway. Initially, T6ODM was characterized as an enzyme catalyzing O-demethylation of thebaine to neopinone and oripavine to morphinone. However, the substrate range of T6ODM was recently expanded to a number of various benzylisoquinoline alkaloids. Here, we present crystal structures of T6ODM in complexes with 2-oxoglutarate (T6ODM:2OG, PDB: 5O9W) and succinate (T6ODM:SIN, PDB: 5O7Y). Both metal and 2OG binding sites display similarity to other proteins from the ODD family, but T6ODM is characterized by an exceptionally large substrate binding cavity, whose volume can partially explain the promiscuity of this enzyme. Moreover, the size of the cavity allows for binding of multiple molecules at once, posing a question about the substrate-driven specificity of the enzyme.
- Jerzy Haber Institute of Catalysis and Surface Chemistry, Polish Academy of Sciences, Niezapominajek 8, PL-30239 Krakow, Poland.
Organizational Affiliation: