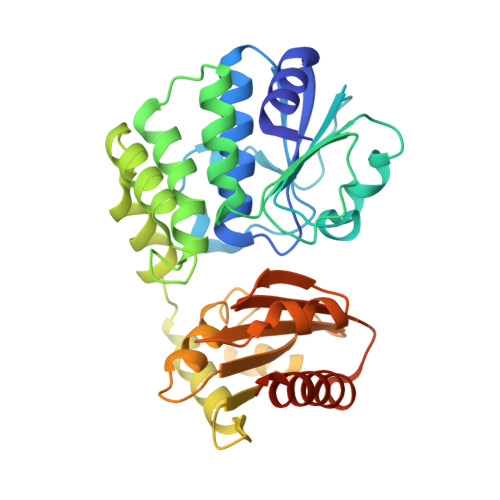
Structural and Biophysical Analysis of the Soluble DHH/DHHA1-Type Phosphodiesterase TM1595 from Thermotoga maritima.
Drexler, D.J., Muller, M., Rojas-Cordova, C.A., Bandera, A.M., Witte, G.(2017) Structure 25: 1887-1897.e4
- PubMed: 29107484 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2017.10.001
- Primary Citation Related Structures:
5O1U, 5O25, 5O4Z, 5O58, 5O70, 5O7F - PubMed Abstract:
The concentration of messenger molecules in bacterial cells needs to be tightly regulated. This can be achieved by either controlling the synthesis rate, degradation, or export by specific transporters, respectively. The regulation of the essential second messenger c-di-AMP is achieved by modulation of the diadenylate cyclase activity as well as by specific phosphodiesterases that hydrolyze c-di-AMP in the cell. We provide here structural and biochemical data on the DHH-type phosphodiesterase TmPDE (TM1595) from Thermotoga maritima. Our analysis shows that TmPDE is preferentially degrading linear dinucleotides, such as 5'-pApA, 5'-pGpG, and 5'-pApG, compared with cyclic dinucleotide substrates. The high-resolution structural data provided here describe all steps of the PDE reaction: the ligand-free enzyme, two substrate-bound states, and three post-reaction states. We can furthermore show that Pde2 from Streptococcus pneumoniae shares both structural features and substrate specificity based on small-angle X-ray scattering data and biochemical assays.
- Gene Center and Department of Biochemistry, Ludwig-Maximilians-Universität München, Feodor-Lynen-Str. 25, 81377 Munich, Germany.
Organizational Affiliation: