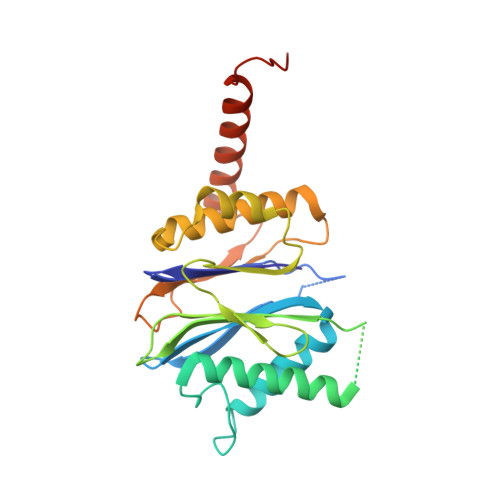
The Y. bercovieri Anbu crystal structure sheds light on the evolution of highly (pseudo)symmetric multimers.
Piasecka, A., Czapinska, H., Vielberg, M.T., Szczepanowski, R.H., Kiefersauer, R., Reed, S., Groll, M., Bochtler, M.(2018) J Mol Biology 430: 611-627
- PubMed: 29258816 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2017.11.016
- Primary Citation Related Structures:
5NYW - PubMed Abstract:
Ancestral β-subunit (Anbu) is homologous to HslV and 20S proteasomes. Based on its phylogenetic distribution and sequence clustering, Anbu has been proposed as the "ancestral" form of proteasomes. Here, we report biochemical data, small-angle X-ray scattering results, negative-stain electron microscopy micrographs and a crystal structure of the Anbu particle from Yersinia bercovieri (YbAnbu). All data are consistent with YbAnbu forming defined 12-14 subunit multimers that differ in shape from both HslV and 20S proteasomes. The crystal structure reveals that YbAnbu subunits form tight dimers, held together in part by the Anbu specific C-terminal helices. These dimers ("protomers") further assemble into a low-rise left-handed staircase. The lock-washer shape of YbAnbu is consistent with the presence of defined multimers, X-ray diffraction data in solution and negative-stain electron microscopy images. The presented structure suggests a possible evolutionary pathway from helical filaments to highly symmetric or pseudosymmetric multimer structures. YbAnbu subunits have the Ntn-hydrolase fold, a putative S 1 pocket and conserved candidate catalytic residues Thr1, Asp17 and Lys32(33). Nevertheless, we did not detect any YbAnbu peptidase or amidase activity. However, we could document orthophosphate production from ATP catalyzed by the ATP-grasp protein encoded in the Y. bercovieri Anbu operon.
- Polish Academy of Sciences, Institute of Biochemistry and Biophysics, Warsaw, Poland; School of Medicine, Cardiff University, Cardiff, United Kingdom.
Organizational Affiliation: