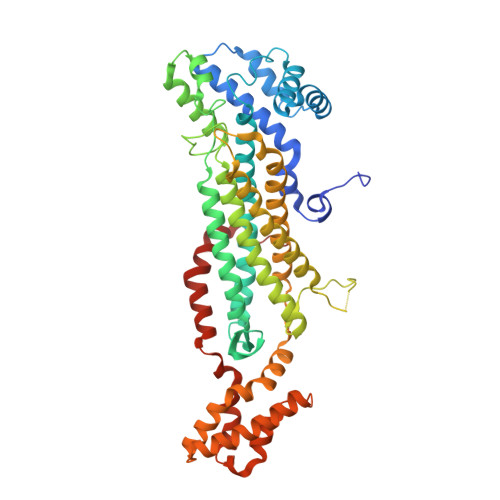
Molecular comparison of Neanderthal and Modern Human adenylosuccinate lyase.
Van Laer, B., Kapp, U., Soler-Lopez, M., Moczulska, K., Paabo, S., Leonard, G., Mueller-Dieckmann, C.(2018) Sci Rep 8: 18008-18008
- PubMed: 30573755 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-018-36195-5
- Primary Citation Related Structures:
5NX8, 5NX9, 5NXA - PubMed Abstract:
The availability of genomic data from extinct homini such as Neanderthals has caused a revolution in palaeontology allowing the identification of modern human-specific protein substitutions. Currently, little is known as to how these substitutions alter the proteins on a molecular level. Here, we investigate adenylosuccinate lyase, a conserved enzyme involved in purine metabolism for which several substitutions in the modern human protein (hADSL) have been described to affect intelligence and behaviour. During evolution, modern humans acquired a specific substitution (Ala429Val) in ADSL distinguishing it from the ancestral variant present in Neanderthals (nADSL). We show here that despite this conservative substitution being solvent exposed and located distant from the active site, there is a difference in thermal stability, but not enzymology or ligand binding between nADSL and hADSL. Substitutions near residue 429 which do not profoundly affect enzymology were previously reported to cause neurological symptoms in humans. This study also reveals that ADSL undergoes conformational changes during catalysis which, together with the crystal structure of a hitherto undetermined product bound conformation, explains the molecular origin of disease for several modern human ADSL mutants.
- Structural Biology Group, European Synchrotron Radiation Facility, CS 40220, F-38043, Grenoble, France.
Organizational Affiliation: