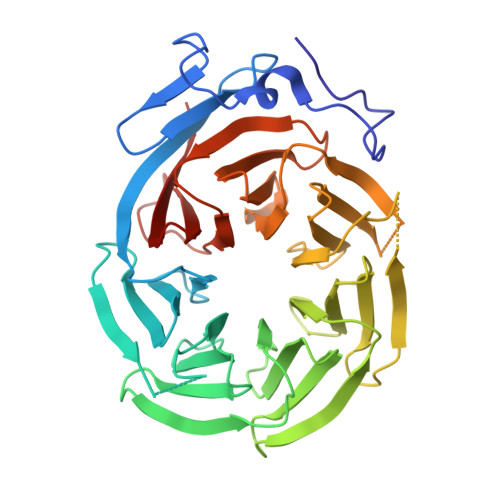
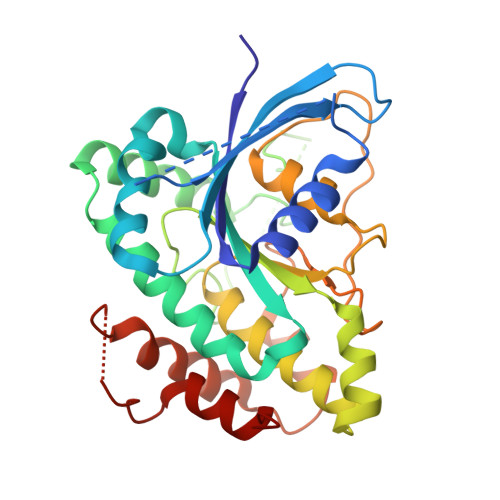
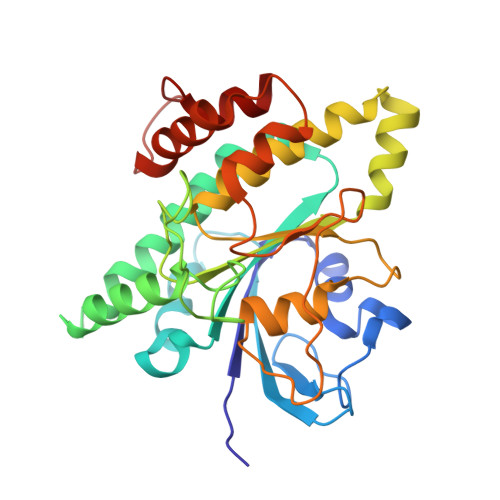
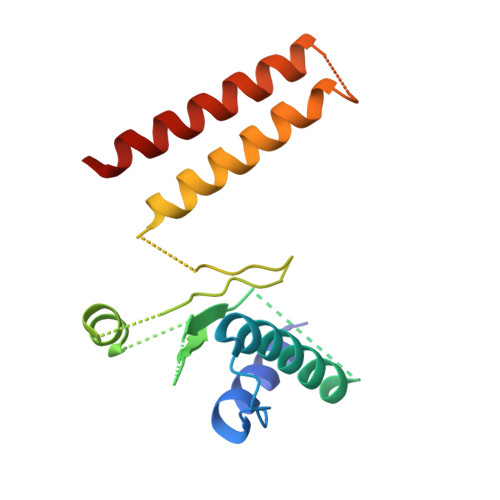
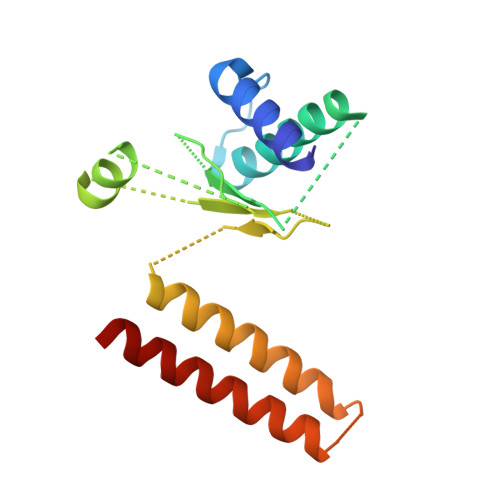
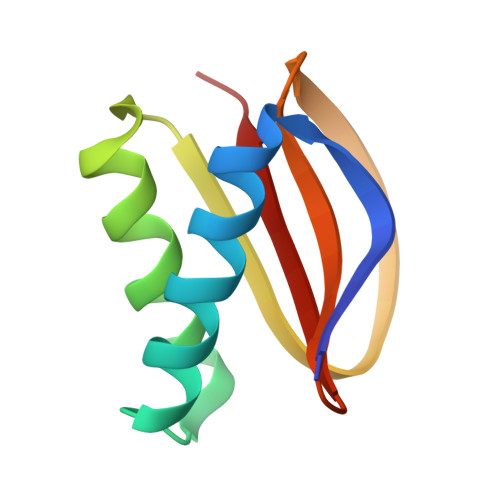
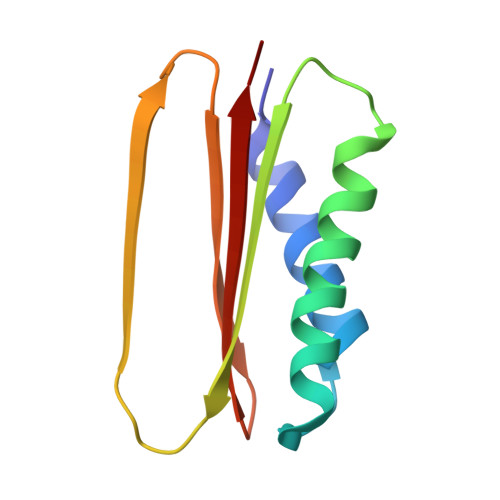
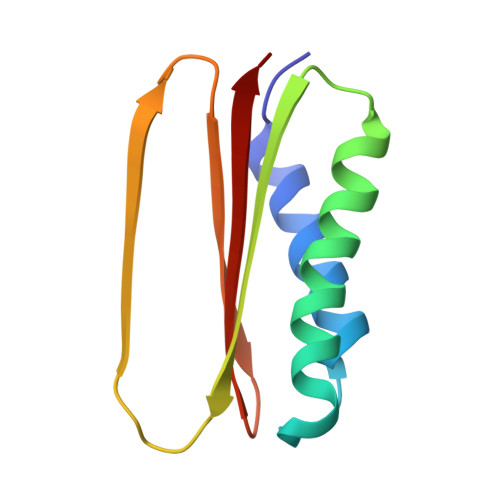
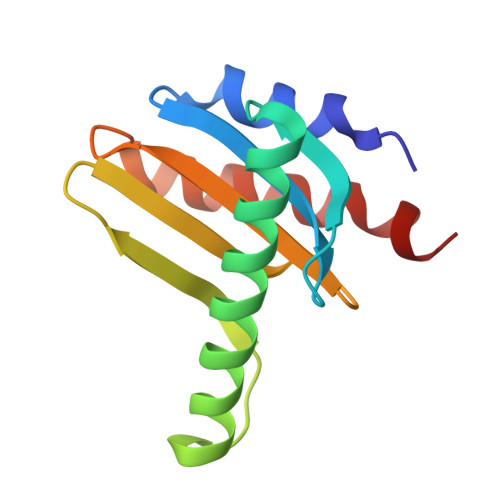
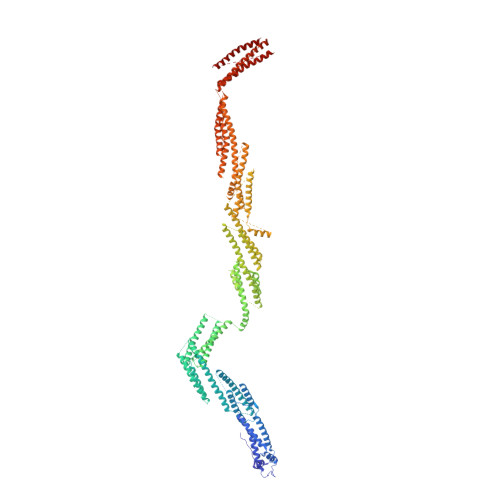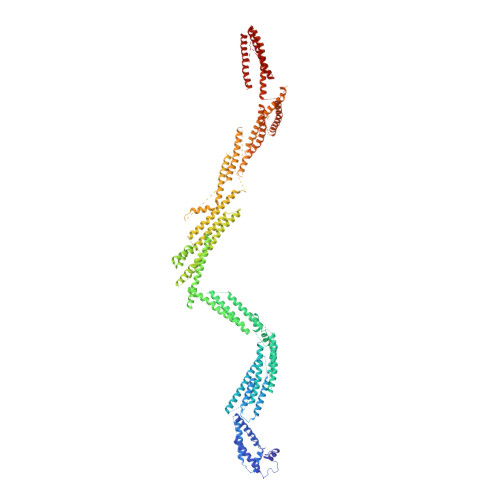Cryo-EM Reveals How Human Cytoplasmic Dynein Is Auto-inhibited and Activated.
Zhang, K., Foster, H.E., Rondelet, A., Lacey, S.E., Bahi-Buisson, N., Bird, A.W., Carter, A.P.(2017) Cell 169: 1303-1314.e18
- PubMed: 28602352 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.cell.2017.05.025
- Primary Citation Related Structures:
5NUG, 5NVS, 5NVU, 5NW4 - PubMed Abstract:
Cytoplasmic dynein-1 binds dynactin and cargo adaptor proteins to form a transport machine capable of long-distance processive movement along microtubules. However, it is unclear why dynein-1 moves poorly on its own or how it is activated by dynactin. Here, we present a cryoelectron microscopy structure of the complete 1.4-megadalton human dynein-1 complex in an inhibited state known as the phi-particle. We reveal the 3D structure of the cargo binding dynein tail and show how self-dimerization of the motor domains locks them in a conformation with low microtubule affinity. Disrupting motor dimerization with structure-based mutagenesis drives dynein-1 into an open form with higher affinity for both microtubules and dynactin. We find the open form is also inhibited for movement and that dynactin relieves this by reorienting the motor domains to interact correctly with microtubules. Our model explains how dynactin binding to the dynein-1 tail directly stimulates its motor activity.
- MRC Laboratory of Molecular Biology, Cambridge CB2 0QH, UK.
Organizational Affiliation: