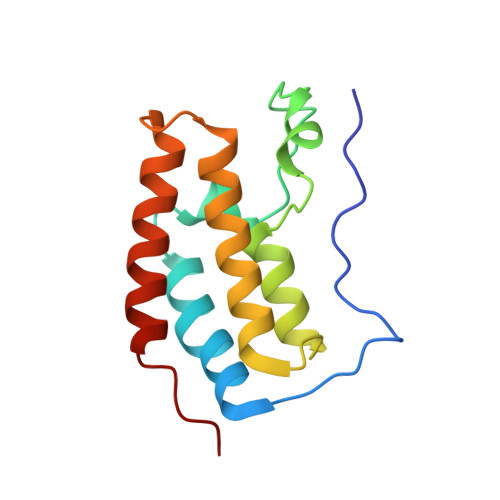
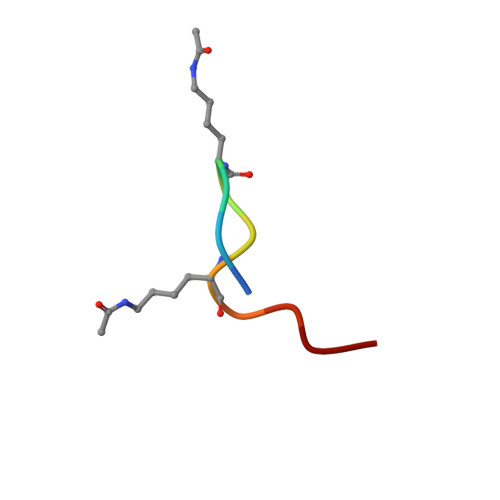
Interactome Rewiring Following Pharmacological Targeting of BET Bromodomains.
Lambert, J.P., Picaud, S., Fujisawa, T., Hou, H., Savitsky, P., Uuskula-Reimand, L., Gupta, G.D., Abdouni, H., Lin, Z.Y., Tucholska, M., Knight, J.D.R., Gonzalez-Badillo, B., St-Denis, N., Newman, J.A., Stucki, M., Pelletier, L., Bandeira, N., Wilson, M.D., Filippakopoulos, P., Gingras, A.C.(2019) Mol Cell 73: 621-638.e17
- PubMed: 30554943 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2018.11.006
- Primary Citation Related Structures:
5NNC, 5NND, 5NNE, 5NNF, 5NNG, 6G0O, 6G0P, 6G0Q, 6G0R, 6G0S - PubMed Abstract:
Targeting bromodomains (BRDs) of the bromo-and-extra-terminal (BET) family offers opportunities for therapeutic intervention in cancer and other diseases. Here, we profile the interactomes of BRD2, BRD3, BRD4, and BRDT following treatment with the pan-BET BRD inhibitor JQ1, revealing broad rewiring of the interaction landscape, with three distinct classes of behavior for the 603 unique interactors identified. A group of proteins associate in a JQ1-sensitive manner with BET BRDs through canonical and new binding modes, while two classes of extra-terminal (ET)-domain binding motifs mediate acetylation-independent interactions. Last, we identify an unexpected increase in several interactions following JQ1 treatment that define negative functions for BRD3 in the regulation of rRNA synthesis and potentially RNAPII-dependent gene expression that result in decreased cell proliferation. Together, our data highlight the contributions of BET protein modules to their interactomes allowing for a better understanding of pharmacological rewiring in response to JQ1.
- Lunenfeld-Tanenbaum Research Institute at Mount Sinai Hospital, Toronto, ON M5G 1X5, Canada.
Organizational Affiliation: