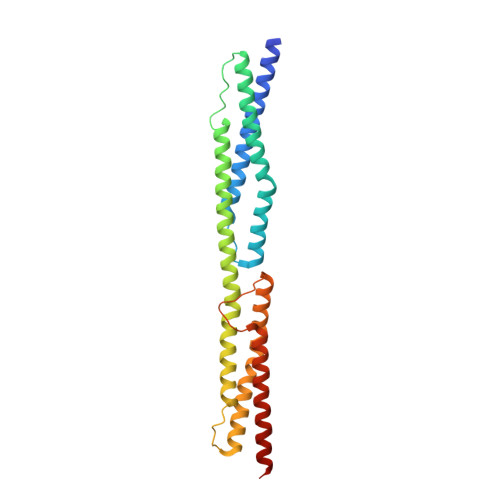
Calcium modulates the domain flexibility and function of an alpha-actinin similar to the ancestral alpha-actinin.
Pinotsis, N., Zielinska, K., Babuta, M., Arolas, J.L., Kostan, J., Khan, M.B., Schreiner, C., Salmazo, A., Ciccarelli, L., Puchinger, M., Gkougkoulia, E.A., Ribeiro Jr., E.A., Marlovits, T.C., Bhattacharya, A., Djinovic-Carugo, K.(2020) Proc Natl Acad Sci U S A 117: 22101-22112
- PubMed: 32848067 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1917269117
- Primary Citation Related Structures:
5NL6, 5NL7, 6SL2, 6SL3, 6SL7 - PubMed Abstract:
The actin cytoskeleton, a dynamic network of actin filaments and associated F-actin-binding proteins, is fundamentally important in eukaryotes. α-Actinins are major F-actin bundlers that are inhibited by Ca 2+ in nonmuscle cells. Here we report the mechanism of Ca 2+ -mediated regulation of Entamoeba histolytica α-actinin-2 ( Eh Actn2) with features expected for the common ancestor of Entamoeba and higher eukaryotic α-actinins. Crystal structures of Ca 2+ -free and Ca 2+ -bound Eh Actn2 reveal a calmodulin-like domain (CaMD) uniquely inserted within the rod domain. Integrative studies reveal an exceptionally high affinity of the Eh Actn2 CaMD for Ca 2+ , binding of which can only be regulated in the presence of physiological concentrations of Mg 2+ Ca 2+ binding triggers an increase in protein multidomain rigidity, reducing conformational flexibility of F-actin-binding domains via interdomain cross-talk and consequently inhibiting F-actin bundling. In vivo studies uncover that Eh Actn2 plays an important role in phagocytic cup formation and might constitute a new drug target for amoebic dysentery.
- Department of Structural and Computational Biology, Max Perutz Labs, University of Vienna, A-1030 Vienna, Austria.
Organizational Affiliation: