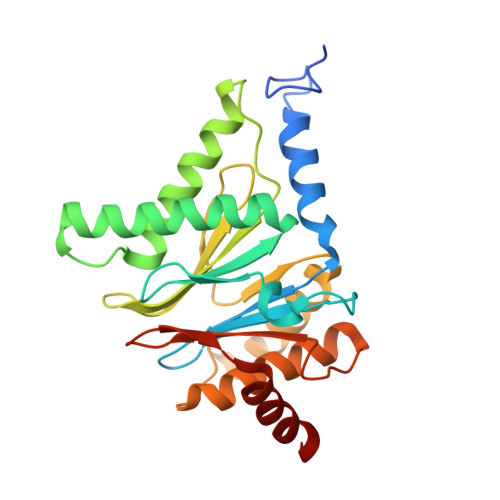
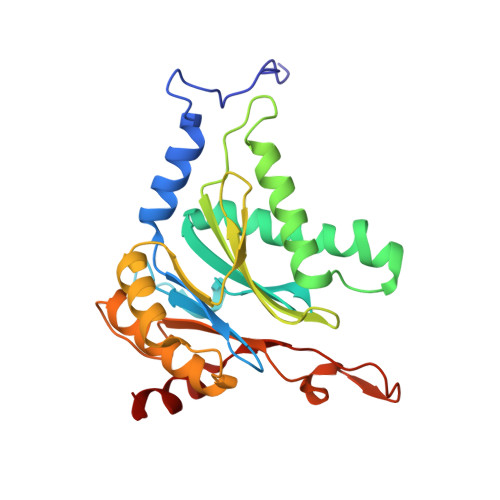
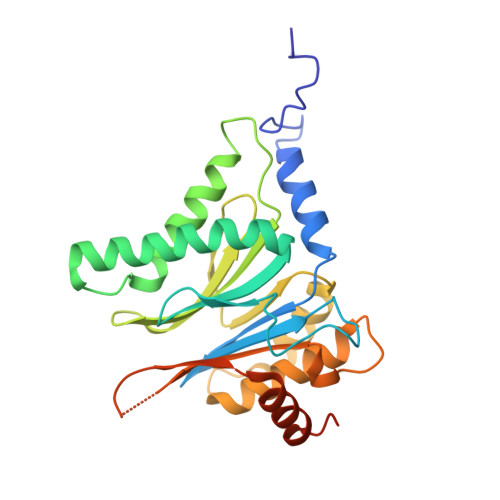
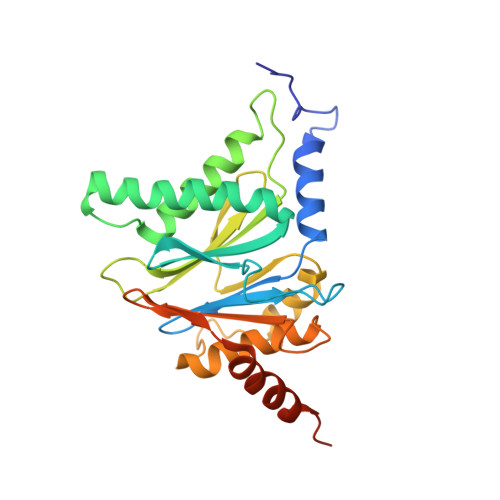
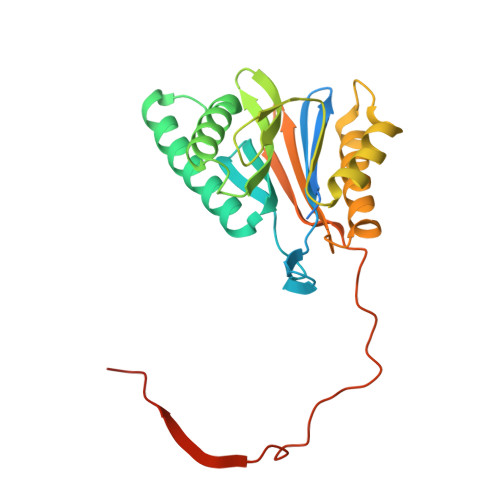
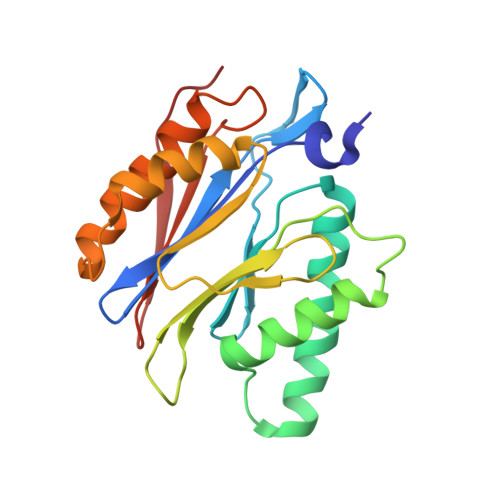
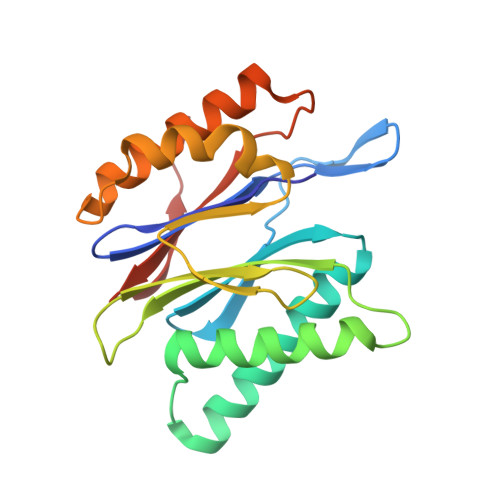
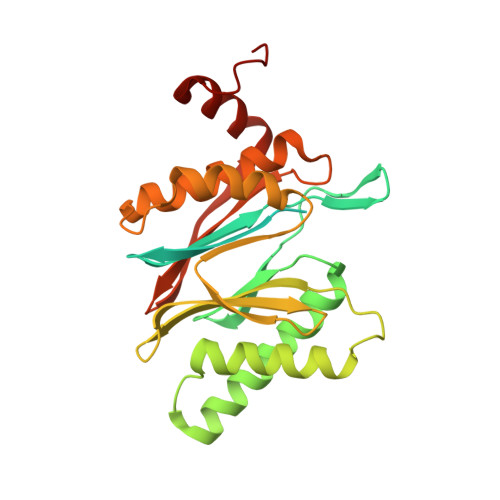
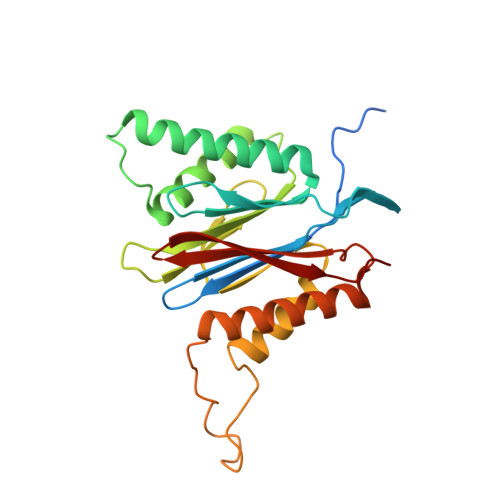
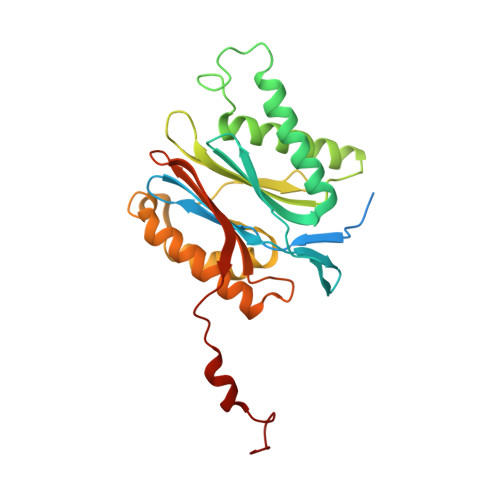
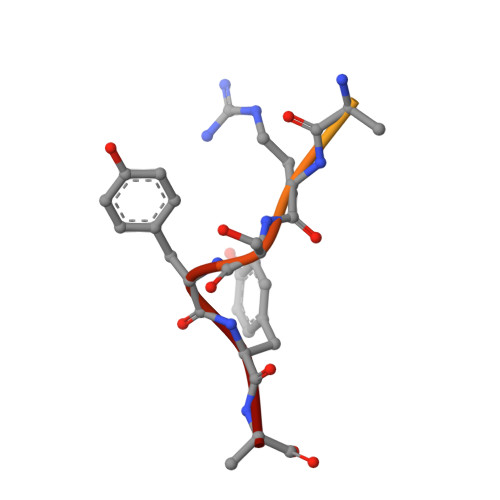
Crystal structure of a low molecular weight activator Blm-pep with yeast 20S proteasome - insights into the enzyme activation mechanism.
Witkowska, J., Gizynska, M., Grudnik, P., Golik, P., Karpowicz, P., Giedon, A., Dubin, G., Jankowska, E.(2017) Sci Rep 7: 6177-6177
- PubMed: 28733623 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-017-05997-4
- Primary Citation Related Structures:
5NIF - PubMed Abstract:
Proteasomes are responsible for protein turnover in eukaryotic cells, degrading short-lived species but also removing improperly folded or oxidatively damaged ones. Dysfunction of a proteasome results in gradual accumulation of misfolded/damaged proteins, leading to their aggregation. It has been postulated that proteasome activators may facilitate removal of such aggregation-prone proteins and thus prevent development of neurodegenerative disorders. However, the discovery of pharmacologically relevant compounds is hindered by insufficient structural understanding of the activation process. In this study we provide a model peptidic activator of human proteasome and analyze the structure-activity relationship within this novel scaffold. The binding mode of the activator at the relevant pocket within the proteasome has been determined by X-ray crystallography. This crystal structure provides an important basis for rational design of pharmacological compounds. Moreover, by providing a novel insight into the proteasome gating mechanism, our results allow the commonly accepted model of proteasome regulation to be revisited.
- Faculty of Chemistry, University of Gdańsk, Wita Stwosza 63, 80-308, Gdańsk, Poland.
Organizational Affiliation: