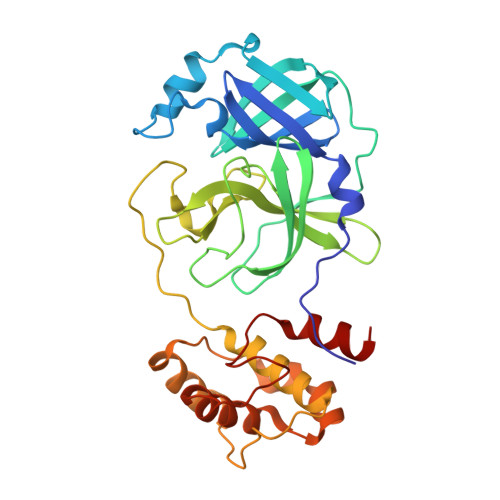
Alpha-ketoamides as broad-spectrum inhibitors of coronavirus and enterovirus replication
Lin, D., Zhang, L., Kusov, Y., Nian, Y., Ma, Q., Wang, J., Leyssen, P., Lanko, K., Neyts, J., de Wilde, A., Snijder, E.J., Liu, H., Hilgenfeld, R.To be published.