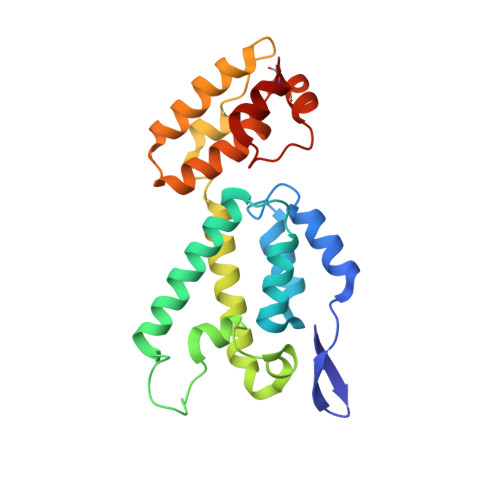
Crystal Structure of the Full-Length Feline Immunodeficiency Virus Capsid Protein Shows an N-Terminal beta-Hairpin in the Absence of N-Terminal Proline.
Folio, C., Sierra, N., Dujardin, M., Alvarez, G., Guillon, C.(2017) Viruses 9
- PubMed: 29120364 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/v9110335
- Primary Citation Related Structures:
5NA2 - PubMed Abstract:
Feline immunodeficiency virus (FIV) is a member of the Retroviridae family. It is the causative agent of an acquired immunodeficiency syndrome (AIDS) in cats and wild felines. Its capsid protein (CA) drives the assembly of the viral particle, which is a critical step in the viral replication cycle. Here, the first atomic structure of full-length FIV CA to 1.67 Å resolution is determined. The crystallized protein exhibits an original tetrameric assembly, composed of dimers which are stabilized by an intermolecular disulfide bridge induced by the crystallogenesis conditions. The FIV CA displays a standard α-helical CA topology with two domains, separated by a linker shorter than other retroviral CAs. The β-hairpin motif at its amino terminal end, which interacts with nucleotides in HIV-1, is unusually long in FIV CA. Interestingly, this functional β-motif is formed in this construct in the absence of the conserved N-terminal proline. The FIV CA exhibits a cis Arg-Pro bond in the CypA-binding loop, which is absent in known structures of lentiviral CAs. This structure represents the first tri-dimensional structure of a functional, full-length FIV CA.
- Equipe Rétrovirus et Biochimie Structurale, Université de Lyon, CNRS, MMSB, UMR 5086 CNRS/Université de Lyon, IBCP, Lyon 69367 CEDEX 07, France. folio.christelle@gmail.com.
Organizational Affiliation: