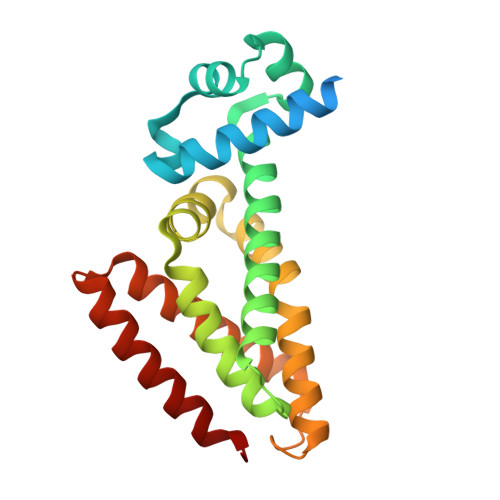
Structural analysis of the interaction between spiroisoxazoline SMARt-420 and the Mycobacterium tuberculosis repressor EthR2.
Wohlkonig, A., Remaut, H., Moune, M., Tanina, A., Meyer, F., Desroses, M., Steyaert, J., Willand, N., Baulard, A.R., Wintjens, R.(2017) Biochem Biophys Res Commun 487: 403-408
- PubMed: 28416386 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2017.04.074
- Primary Citation Related Structures:
5N7O - PubMed Abstract:
Inhibition of transcriptional regulators of bacterial pathogens with the aim of reprogramming their metabolism to modify their antibiotic susceptibility constitutes a promising therapeutic strategy. One example is the bio-activation of the anti-tubercular pro-drug ethionamide, which activity could be enhanced by inhibiting the transcriptional repressor EthR. Recently, we discovered that inhibition of a second transcriptional repressor, EthR2, leads to the awakening of a new ethionamide bio-activation pathway. The x-ray structure of EthR2 was solved at 2.3 Å resolution in complex with a compound called SMARt-420 (Small Molecule Aborting Resistance). Detailed comparison and structural analysis revealed interesting insights for the upcoming structure-based design of EthR2 inhibitors as an alternative to revert ethionamide resistance in Mycobacterium tuberculosis.
- Center for Structural Biology, Vlaams Instituut voor Biotechnology (VIB), Pleinlaan 2, B-1050 Brussels, Belgium; Structural Biology Brussels, Vrije Universiteit Brussel (VUB), Pleinlaan 2, B-1050 Brussels, Belgium.
Organizational Affiliation: