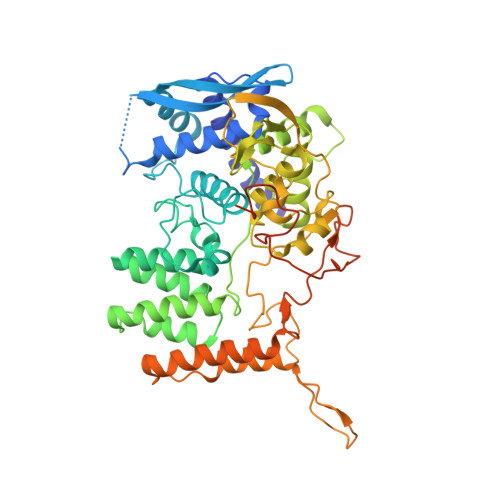
The structure of the nucleoprotein of Influenza D shows that all Orthomyxoviridae nucleoproteins have a similar NPCORE, with or without a NPTAILfor nuclear transport.
Donchet, A., Oliva, J., Labaronne, A., Tengo, L., Miloudi, M., C A Gerard, F., Mas, C., Schoehn, G., W H Ruigrok, R., Ducatez, M., Crepin, T.(2019) Sci Rep 9: 600-600
- PubMed: 30679709 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-018-37306-y
- Primary Citation Related Structures:
5N2U - PubMed Abstract:
This paper focuses on the nucleoprotein (NP) of the newly identified member of the Orthomyxoviridae family, Influenza D virus. To date several X-ray structures of NP of Influenza A (A/NP) and B (B/NP) viruses and of infectious salmon anemia (ISA/NP) virus have been solved. Here we purified, characterized and solved the X-ray structure of the tetrameric D/NP at 2.4 Å resolution. The crystal structure of its core is similar to NP of other Influenza viruses. However, unlike A/NP and B/NP which possess a flexible amino-terminal tail containing nuclear localization signals (NLS) for their nuclear import, D/NP possesses a carboxy-terminal tail (D/NP TAIL ). We show that D/NP TAIL harbors a bipartite NLS and designed C-terminal truncated mutants to demonstrate the role of D/NP TAIL for nuclear transport.
- Institut de Biologie Structurale (IBS), Univ. Grenoble Alpes, CEA, CNRS, 38044, Grenoble, France.
Organizational Affiliation: