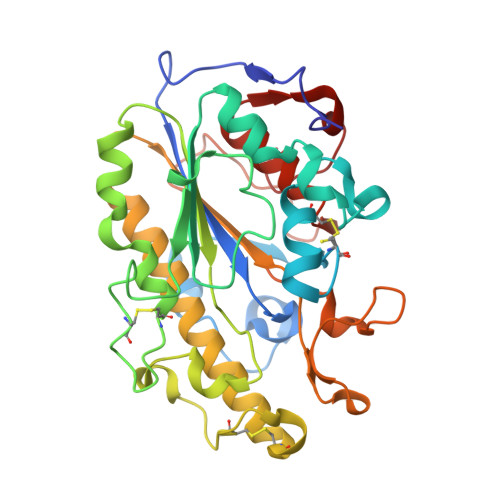
1.12 angstrom resolution crystal structure of the catalytic domain of the plasmid-mediated colistin resistance determinant MCR-2.
Coates, K., Walsh, T.R., Spencer, J., Hinchliffe, P.(2017) Acta Crystallogr F Struct Biol Commun 73: 443-449
- PubMed: 28777086 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X17009669
- Primary Citation Related Structures:
5MX9 - PubMed Abstract:
MCR-2 confers resistance to colistin, a `last-line' antibiotic against extensively resistant Gram-negative pathogens. It is a plasmid-encoded phosphoethanolamine transferase that is closely related to MCR-1. To understand the diversity in the MCR family, the 1.12 Å resolution crystal structure of the catalytic domain of MCR-2 was determined. Variable amino acids are located distant from both the di-zinc active site and the membrane-proximal face. The exceptionally high resolution will provide an accurate starting model for further mechanistic studies.
- School of Cellular and Molecular Medicine, University of Bristol, Bristol BS8 1TD, England.
Organizational Affiliation: