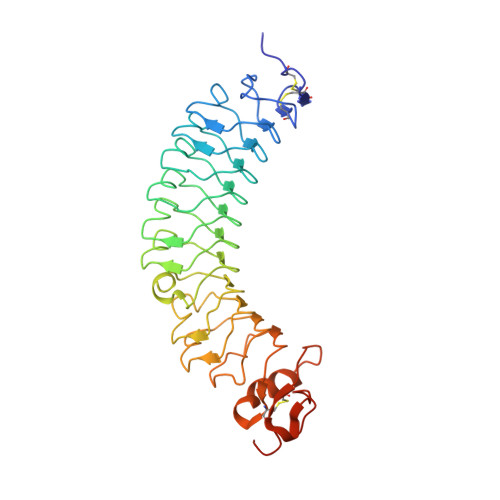
Structural and functional analysis of two small leucine-rich repeat proteoglycans, fibromodulin and chondroadherin.
Paracuellos, P., Kalamajski, S., Bonna, A., Bihan, D., Farndale, R.W., Hohenester, E.(2017) Matrix Biol 63: 106-116
- PubMed: 28215822 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.matbio.2017.02.002
- Primary Citation Related Structures:
5MX0, 5MX1 - PubMed Abstract:
The small leucine-rich proteoglycans (SLRPs) are important regulators of extracellular matrix assembly and cell signalling. We have determined crystal structures at ~2.2Å resolution of human fibromodulin and chondroadherin, two collagen-binding SLRPs. Their overall fold is similar to that of the prototypical SLRP, decorin, but unlike decorin neither fibromodulin nor chondroadherin forms a stable dimer. A previously identified binding site for integrin α2β1 maps to an α-helix in the C-terminal cap region of chondroadherin. Interrogation of the Collagen Toolkits revealed a unique binding site for chondroadherin in collagen II, and no binding to collagen III. A triple-helical peptide containing the sequence GAOGPSGFQGLOGPOGPO (O is hydroxyproline) forms a stable complex with chondroadherin in solution. In fibrillar collagen I and II, this sequence is aligned with the collagen cross-linking site KGHR, suggesting a role for chondroadherin in cross-linking.
- Department of Life Sciences, Imperial College London, London, United Kingdom.
Organizational Affiliation: