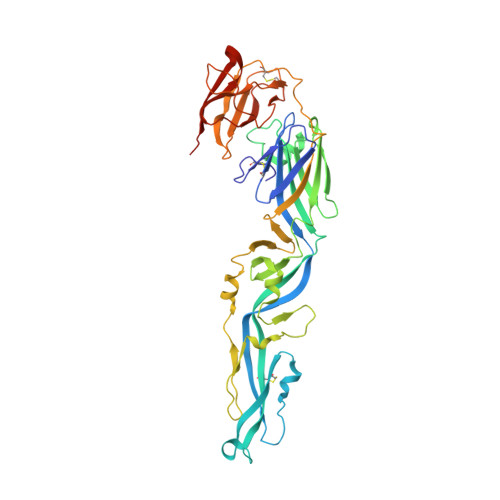
The structure differences of Japanese encephalitis virus SA14 and SA14-14-2 E proteins elucidate the virulence attenuation mechanism.
Liu, X., Zhao, X., Na, R., Li, L., Warkentin, E., Witt, J., Lu, X., Yu, Y., Wei, Y., Peng, G., Li, Y., Wang, J.(2019) Protein Cell 10: 149-153
- PubMed: 29752689 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1007/s13238-018-0551-6
- Primary Citation Related Structures:
5MV1, 5MV2 - National Institutes for Food and Drug Control, Beijing, 102609, China.
Organizational Affiliation: