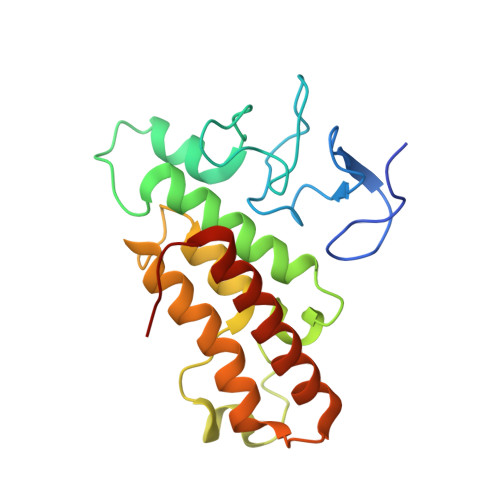
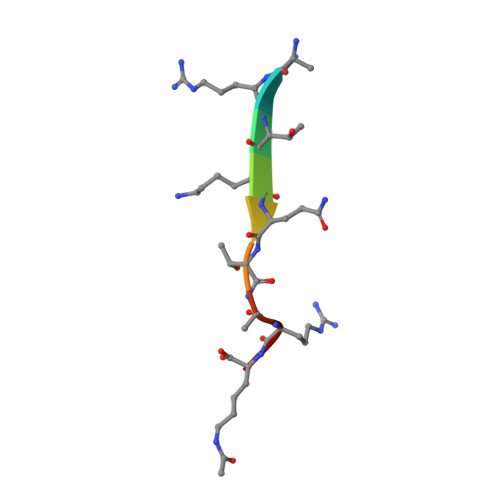
Crystal structure of TRIM33 PHD-Bromodomain isoform B in complex with H3K9ac histone peptide
Tallant, C., Savitsky, P., Fedorov, O., Nunez-Alonso, G., Siejka, P., Krojer, T., Williams, E., Srikannathasan, V., von Delft, F., Arrowsmith, C.H., Edwards, A.M., Bountra, C., Muller, S., Knapp, S.To be published.