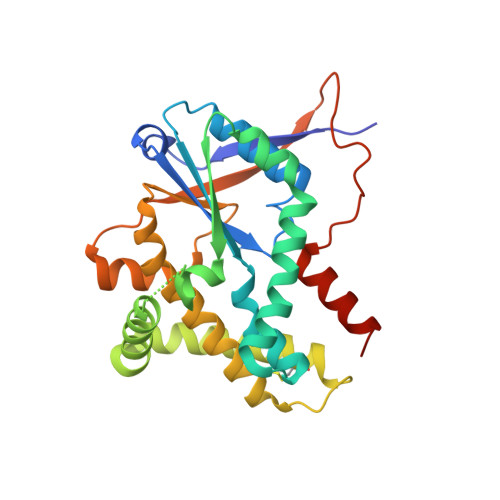
Structural basis for partition of the cyclodipeptide synthases into two subfamilies.
Bourgeois, G., Seguin, J., Babin, M., Belin, P., Moutiez, M., Mechulam, Y., Gondry, M., Schmitt, E.(2018) J Struct Biol 203: 17-26
- PubMed: 29505829 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2018.03.001
- Primary Citation Related Structures:
5MLP, 5MLQ, 5OCD, 6EZ3 - PubMed Abstract:
Cyclodipeptide synthases (CDPSs) use two aminoacyl-tRNAs to catalyze the formation of two peptide bonds leading to cyclodipeptides that can be further used for the synthesis of diketopiperazines. It was shown that CDPSs fall into two subfamilies, NYH and XYP, characterized by the presence of specific sequence signatures. However, current understanding of CDPSs only comes from studies of enzymes from the NYH subfamily. The present study reveals the crystal structures of three CDPSs from the XYP subfamily. Comparison of the XYP and NYH enzymes shows that the two subfamilies mainly differ in the first half of their Rossmann fold. This gives a structural basis for the partition of CDPSs into two subfamilies. Despite these differences, the catalytic residues adopt similar positioning regardless of the subfamily suggesting that the XYP and NYH motifs correspond to two structural solutions to facilitate the reactivity of the catalytic serine residue.
- Laboratoire de Biochimie, Ecole polytechnique, CNRS, Université Paris-Saclay, 91128 Palaiseau cedex, France.
Organizational Affiliation: