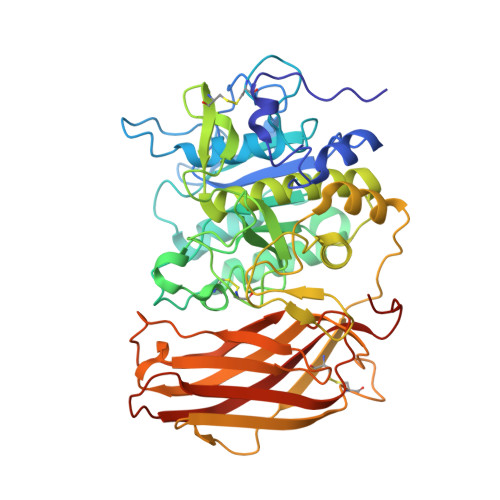
Structural Studies Revealed Active Site Distortions of Human Furin by a Small Molecule Inhibitor.
Dahms, S.O., Jiao, G.S., Than, M.E.(2017) ACS Chem Biol 12: 1211-1216
- PubMed: 28402100 Search on PubMed
- DOI: https://doi.org/10.1021/acschembio.6b01110
- Primary Citation Related Structures:
5MIM - PubMed Abstract:
Proprotein convertases (PCs) represent highly selective serine proteases that activate their substrates upon proteolytic cleavage. Their inhibition is a promising strategy for the treatment of several pathologies including cancer, atherosclerosis, hypercholesterolaemia, and infectious diseases. Here, we present the first experimental complex of furin with a non-substrate-like small molecule inhibitor, and the X-ray structure of the enzyme complexed to the small molecule inhibitor 1 at 1.9 Å resolution. Two molecules of inhibitor 1 were found to interact with furin. One is anchored at the S4 pocket of the enzyme and interferes directly with the conformation and function of the catalytic triade; the other molecule shows weaker binding and interacts with a distant, less conserved region of furin. The observed binding modes represent a new inhibition strategy of furin and imply the possibility to attain specificity among the PCs providing an innovative starting point of structure guided inhibitor development for furin.
- Protein Crystallography Group, Leibniz Institute on Aging - Fritz Lipmann Institute (FLI) , Beutenbergstr. 11, 07745 Jena, Germany.
Organizational Affiliation: