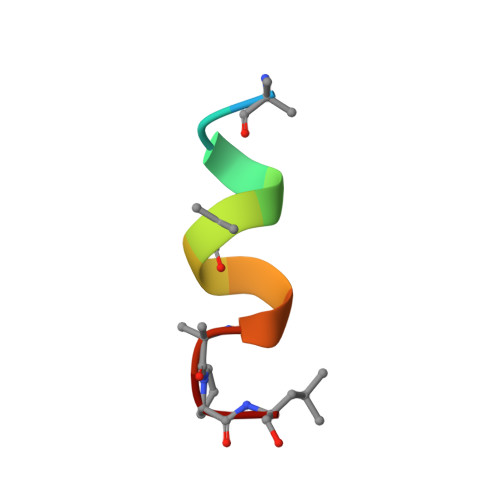11-mer peptaibol Harzianin HK-VI: conformational and biological analysis
Kara, S., Afonin, S., Zamora-Carreras, H., Bordessa, A., Doan, V., Takeshita, N., Grage, S.L., Chaume, G., Jimenez, M.A., Fischer, R., Bruix, M., Brigaud, T., Ulrich, A.S.To be published.